INTRODUCTION
Aruncus aethusifolius (H. Lév.) Nakai (Rosaceae; Spiraeoideae; Spiraeeae) is an endemic species of Korea, distributed only on Jejudo Island. Plants of A. aethusifolius are found at the outcrops of volcanic rocks at elevations ranging from 1,500 to 1,950 m, and they also grow in moist parts of lowlands on the island (Yun et al., 2017). Aruncus aethusifolius is a small perennial herb up to 30 cm tall with estipulate, pinnately compound leaves; compact panicles with many small, unisexual or rarely bisexual flowers; and erect pedicels and follicles. It is morphologically similar to A. dioicus var. kamtschaticus (Maxim.) H. Hara and A. dioicus var. astilboides (Maxim.) H. Hara, but easily distinguished from these by having highly dissected leaflets (Nakai, 1912; Lee, 1980; Lee, 2006; Yun et al., 2017). Aruncus aethusifolius is widely cultivated as an ornamental, known as dwarf goat’s beard, for its attractive inflorescence arising above the foliage mound and because it requires little care to grow. As a part of biogeographic and population genetic studies of the species, the complete chloroplast genome was determined to understand the origin of this economically important plant.
MATERIALS AND METHODS
The sample was collected on Georinjokeun Oreum on Jejudo Island, Korea (33º21′8.20″N, 126º39′48.50″E). A specimen was deposited at the Daejeon University Herbarium (TUT) under the voucher number Suh 8681. It is locally common. No permission is required for the collection. Total DNA was extracted from 100 mg of fresh leaves using a DNeasy Plant Mini Kit (QIAGEN, Hilden, Germany), and visualized in 1% agarose gel. An amount of 1.2 micrograms of DNA, measured with an Invitrogen Qubit fluorometer, was used to construct the library. The sequencing library was constructed using an Illumina TruSeq Nano DNA Library Preparation Kit (Illumina, San Diego, CA, USA) following the manufacturer’s recommendations with approximately 350-bp DNA fragments. 1.99-Gbp raw sequences were obtained using HiSeqX at Macrogen Inc. (Seoul, Korea), and were filtered by Trimmomatic v0.33 (Bolger et al., 2014). The chloroplast genome was de novo assembled with Velvet v1.2.10 (Zerbino and Birney, 2008), and gaps were closed using GapCloser v1.12 (Zhao et al., 2011). The genome sequence was confirmed by aligning all raw reads against the assembled genome using BWA v0.7.17 and SAMtools v1.9 (Li, 2013; Li et al., 2009).
All processes were conducted in the Genome Information System utilized in previous studies (Park et al., 2019c; Baek et al., 2021; Park et al., 2021). Geneious Prime v2020.2.4 (Biomatters Ltd., Auckland, New Zealand) was used for annotation based on the A. dioicus var. kamtschaticus chloroplast (MW115132) (Suh et al., 2021). A circular map of A. aethusifolius chloroplast genome was drawn using OGDRAW v1.31 (Greiner et al., 2019). Large single-copy (LSC), small single-copy (SSC), and inverted repeat (IR) regions were determined by bl2seq (Tatusova and Madden, 1999), which provides a dot plot to show exact IR regions.
Sixteen whole chloroplast genomes to represent the major lineages of the tribe Spiraeeae (Potter et al., 2007) were included in phylogenetic analyses using the maximum likelihood (ML), neighbor-joining, and Bayesian inference (BI) methods. Heuristic searches were employed to reconstruct the ML phylogenetic tree using MEGA X (Kumar et al., 2018) with the following options: nearest-neighbor interchange branch swapping, the Tamura-Nei model, uniform rates among sites, and default values for other options. Bootstrap analyses with 1,000 pseudoreplicates were also conducted. The BI tree was constructed using MrBayes v3.2.6 (Ronquist et al., 2012). The GTR model with gamma rates was used, determined in jModelTest v2.1.5 (Darriba et al., 2012). A Markov-chain Monte Carlo algorithm was employed for 1,000,000 generations, sampling trees every 200 generations, with four chains running simultaneously.
RESULTS AND DISCUSSION
The chloroplast genome of A. aethusifolius (GenBank accession: MZ882398) (Fig. 1) is 157,217 bp long (GC ratio is 36.5%) and has four subregions consisting of 85,207 bp of LSC (34.3%) and 19,222 bp of SSC (30.0%) regions separated by 26,394 bp of IR regions (42.4%). It contains 131 genes (86 protein-coding genes, eight rRNAs, and 37 tRNAs); 19 genes (eight protein-coding genes, four rRNAs, and seven tRNAs) are duplicated in IR regions (Fig. 1). The chloroplast genome sequence can be accessed via accession number MZ882398 and SRA numbers are PRJNA722681, SAMN18789871, and SRR14267045.
Interspecific variations between A. aethusifolius and A. dioicus var. kamtschaticus were investigated with the same method used in the previous studies (Kim et al., 2021; Lee and Park, 2021; Yoo et al., 2021). In total, there were 206 single nucleotide polymorphisms and 62 regions of insertion/deletion (INDEL regions, 907 bp in total). Remarkably, a 687-bp deletion was found in an intergenic region between trnT-GGU and psbD between the chloroplast genomes of A. dioicus var. kamtschaticus and A. aethusifolius, which contributes to the major difference in the genome size between the two (642 bp). These kinds of long INDELs were also found in the chloroplast genomes of Arabidopsis thaliana L. (378 bp) (Park et al., 2020a) and Coffea arabica L. (84 bp) (Park et al., 2019b) and mitochondrial genomes of Liriodendron tulipifera L. (1,980 bp) (Park et al., 2019a) and Rosa rugosa Thunb. ex Murray (607 bp) (Park et al., 2020b). Moreover, two 26-bp insertions found in the SSC region and 17 INDEL regions, of which length is above 5-bp were also identified. These interspecific variations are similar to the chloroplast genomes of four Viburnum species (Park et al., 2020c).
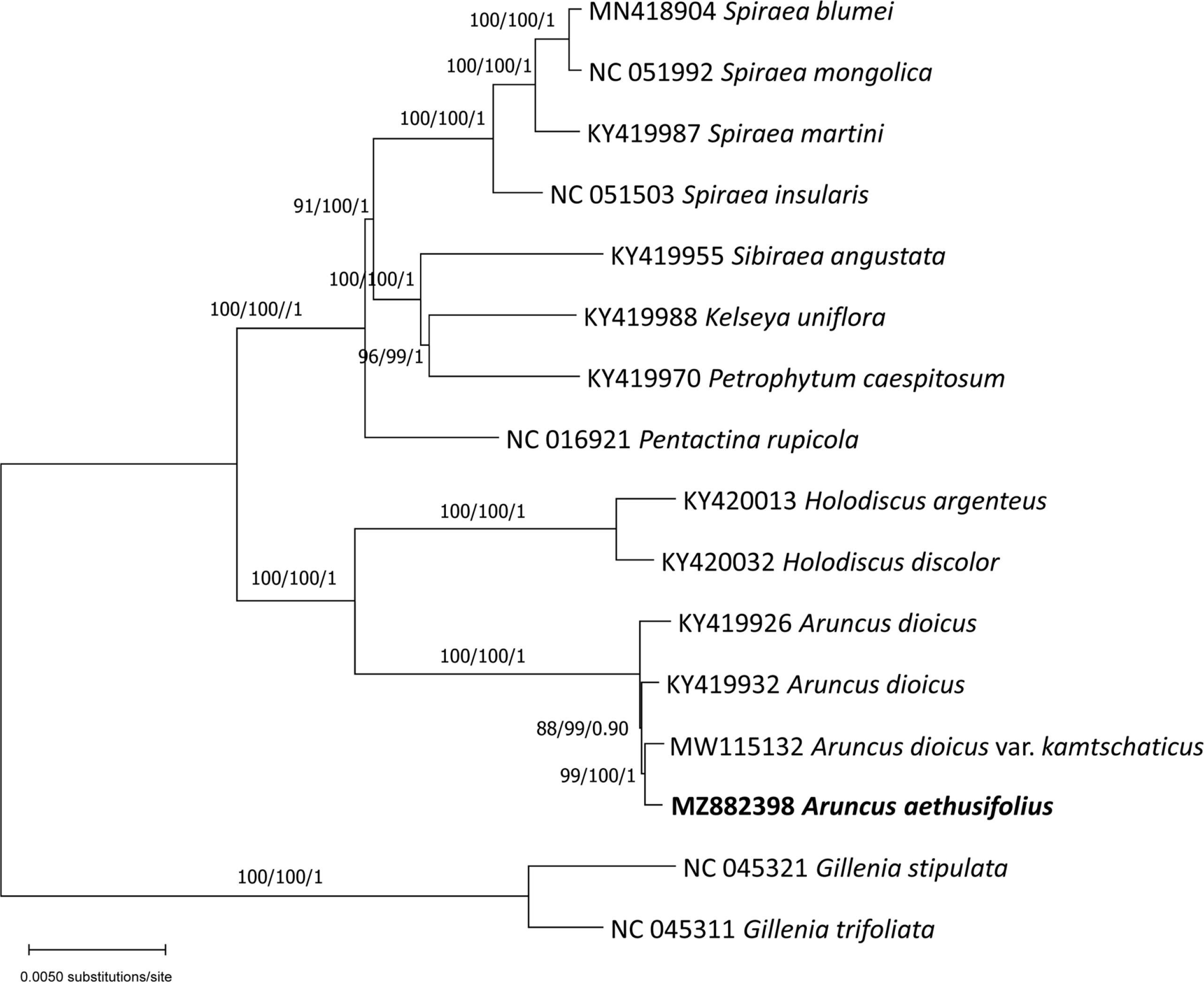
The phylogenetic tree showed that A. aethusifolius formed a strongly supported clade of Aruncus with A. dioicus and A. dioicus var. kamtschaticus, sister to Holodiscus (K. Koch) Maxim. (Fig. 2). The results of phylogenetic analyses are consistent with previous data, in which the clade of Aruncus and monotypic Leutkea Bong. is sister to Holodiscus (Potter et al., 2007; Zhang et al., 2017). The precise phylogenetic position of A. aethusifolius within Aruncus is important to gain insights into the origin of the species. But, chloroplast genome data of other varieties of A. dioicus (Hara, 1955) could not be included in this analysis because they are unavailable. Aruncus dioicus var. astilboides, a rare species in northern Honshu, Japan, is of particular interest because erect follicles found in A. aethusifolius are also characteristics of the Japanese variety. The information of organisms for the two genome data of A. dioicus from GenBank (GenBank accessions: KY419926 and KY419932) were not provided to the varietal level. Further studies with more taxa are needed to infer the origin of A. aethusifolius and to understand the biogeographic history of Aruncus. This is the first complete chloroplast genome sequence of A. aethusifolius, which will be helpful to develop species-specific markers.