/home/virtual/kjpt/journal//../xmls/kjpt-52-2-102.xml
CHOI, KIM, BYUN, GANG, LEE, MYEONG, SO, and JANG: A study of the chromosome number and genome size of the rare species Rhododendron keiskei var. hypoglaucum in Korea
Abstract
Rhododendron keiskei var. hypoglaucum (Ericaceae) was recently reported in Korea, with a disjunct distribution on the southern islands of the Korean Peninsula. Although chromosome numbers and ploidy variations are important traits in angiosperms, gaining a clear understanding the cytological features of Rhododendron has been hampered by the small size of its chromosomes. We herein report the chromosome number, karyotype structure, and genome size of R. keiskei var. hypoglaucum for the first time. The chromosome number of the investigated plants was 2n = 26 with x = 13 as the base chromosome number, which is the one of the frequently encountered base chromosome numbers in Rhododendron. The karyotype of R. keiskei var. hypoglaucum is composed of metacentric and submetacentric chromosomes similar in length, which ranged from 1.39 to 2.40 μm. The DNA 1C-value in all examined accessions was small, ranging from 0.63 to 0.65 pg, further supporting the stable genome size in Rhododendron. These comprehensive cytological results provide a framework for detailed molecular, cytogenetic, and phylogenomic analyses that can be used to interpret the slow species diversification rate in Rhododendron.
Keywords: Chromosome number, flow cytometry, genome size, karyotype, Rhododendron keiskei var. hypoglaucum
INTRODUCTION
The changes in chromosome number such as polyploidy and dysploidy play an important role in plant evolution, diversification, and eventually speciation ( Lysak et al., 2006; Schubert, 2007; Soltis and Soltis, 2009). To date, chromosome numbers have been reported for approximately 25–30% of flowering plants ( Weiss-Schneeweiss and Schneeweiss, 2013; Rice et al., 2015) and, along with molecular phylogenetic data ( Pessoa et al., 2022) and genome size data ( Choi et al., 2020; Mitrenina et al., 2021; Greimler et al., 2022), are still widely used in systematics. The genome size (1C value) may vary considerably among closely related species, but is usually consistent within most species ( Bennett et al., 2000; Greilhuber, 2005). Thus, it has important systematic implications ( McCann et al., 2018; Choi et al., 2019, 2020), although significant variations in genome size have been observed among populations within the same species ( Jang et al., 2018; Agudo et al., 2019; Becher et al., 2021). Classical karyotyping methods using Feulgen staining are challenging in highly polyploid species of Cyperaceae, Juncaceae and Ericaceae due to small size of the chromosomes, which hinders chromosomal determination in polyploids and aneuploids ( Chung and Im, 2020; Chung and Chung, 2021; Choi et al., 2022; Redpath et al., 2022). Flow cytometry is an alternative method for estimating genome size and ploidy level ( Choi et al., 2021; Sliwinska et al., 2021; Temsch et al., 2021).
Rhododendron is the largest genus of the family Ericaceae, comprising about 1,000 species. The genus includes perennial woody plants distributed in Asia, Europe, North America, and Australia ( Fang et al., 2005). Along with one recently recorded species ( R. keiskei Miq. var. hypoglaucum Sutô & Suzuki; Yang et al., 2015), 12 taxa are currently recognized in Korea ( Chang, 2007). The base chromosome number of Rhododendron is x = 13, and different ploidy levels (2 x, 4 x, and 6 x) have been documented ( Sax, 1930; Rice et al., 2015). Chromosome number and variation in genome size have been used in the systematics of Ericaceae ( Khan et al., 2021). Information on chromosome number and C-values in Korean Rhododendron taxa is scarce ( Pellicer and Leitch, 2020; Rice et al., 2015). At present, all Korean species are reported only as diploids ( Appendix 1), and detailed karyotype analyses of these taxa are lacking. In our study, we investigated the chromosome number, karyotype, and genome size of the recently recorded species in Korea, R. keiskei var. hypoglaucum, for the first time.
MATERIALS AND METHODS
Six individuals from the recently recorded species Rhododendron keiskei var. hypoglaucum were collected from a natural population in Korea, and cultivated in the Plant Conservation Center (Korea National Park Research Institute) ( Fig. 1, Table 1). Actively growing root tips from the cultivated plants were pretreated with 0.2 mM 8-hydroxyquinoline solution for 2.5 h at room temperature and then for another 2.5 h at 4°C before the samples were fixed in ethanol: acetic acid (3:1, v/v). Chromosome numbers and karyotypes were determined using the standard Feulgen staining technique as described by Choi et al. (2019) and Ikeda et al. (2021). The chromosomes of the investigated taxon are very small. Therefore, to improve karyotype resolution, the cells were additionally digested with enzymes as described by Jang et al. (2016). Briefly, meristematic root cells were digested at 37°C for 2.5 h with 1% cellulase Onozuka (Serva, Heidelberg, Germany), 1% cytohelicase (Sigma-Aldrich, Vienna, Austria), and 1% pectolyase (Sigma-Aldrich). Squash preparations were made in a drop of 60% acetic acid, and prepared slides were snap frozen in liquid nitrogen and cover slips were removed. All prepared slides were stained with 2 ng/μL 4′,6-diamidino- 2-2phenylindole (DAPI) dissolved in the mounting antifade medium Vectashield (Vector Laboratories, Burlingame, CA, USA) and examined under a fluorescence microscope (BX-51, Olympus, Tokyo, Japan). Representative karyotypes were selected, cut out, and arranged in Adobe Photoshop CS6. Chromosome size was measured using Micromeasure, ver. 3.3 ( http://www.colostate.edu/Depts/Boology/MicroMeasure/) as described by Jang et al. (2013). The genome size of the six individuals of R. keiskei var. hypoglaucum was measured with flow cytometry (Sysmex CyFlow Cytometer, Sysmex Partec GmbH, Görlitz, Germany) performed on leaf materials. Fresh tissue from the cultivated plants growing in the Plant Conservation Center were used with Solanum pseudocapsicum L. (1C = 1.30 pg) ( Temsch et al., 2010) as an internal standard. The detailed methodology used for the genome size measurement was described by Choi et al. (2021).
RESULTS AND DISCUSSION
Chromosome number and karyotype
The chromosome number determined for Rhododendron keiskei var. hypoglaucum is reported here for the first time ( Table 1). Additional information on earlier chromosome reports for Korean Rhododendron taxa is given in Appendix 1. The base chromosome number of R. keiskei var. hypoglaucum was inferred to be x = 13 (2 n = 2 x = 26) ( Fig. 2), in agreement with previous reports on chromosome numbers in other Rhododendron species ( Gurzenkov, 1973; Murin et al., 1984; Stepanov, 1994; Atkinson et al., 2000). The previously reported chromosome numbers in 227 Rhododendron species included three different ploidy levels (i.e., 2 x, 4 x, and 6 x) ( Rice et al., 2015). Such relatively stable ploidy states and variations in chromosome number have been often documented in woody plants ( Stebbins, 1938; Otto and Whitton, 2000), although little experimental research beyond comparative surveys on the chromosome number variations was conducted in angiosperms ( Husband et al., 2013). Cut-out karyotypes were prepared for one individual of R. keiskei var. hypoglaucum to protect the taxa from the natural population ( Figs. 1, 2). The karyotype of R. keiskei var. hypoglaucum is characterized by an asymmetry index (AsI) of 57.4%, predominantly metacentric and submetacentric chromosomes ( Fig. 2B, C), the chromosome length in the range of 1.39 to 2.40 μm, and the haploid karyotype length of 25.47 μm ( Fig. 2C). Similar to other Rhododendron taxa ( Sax, 1930; Atkinson et al., 2000), identification of individual chromosome pairs was challenging. Due to this reason, only one pair of nucleolar organizer region was documented in the R. keiskei var. hypoglaucum karyotype ( Fig. 2A), in accord with an earlier study ( Atkinson et al., 2000).
Nuclear DNA content
We report for the first time the genome size of Rhododendron keiskei var. hypoglaucum ( Table 1); flow histograms are shown in Fig. 3A. Genome size (1C) measured for six individuals ranged from 0.63 to 0.65 pg ( Fig. 3B), which falls within the range of that reported in other species of the genus (e.g., 1C = 0.483–2.777 pg, but chromosome number unknown) ( Khan et al., 2021). The genome size of R. keiskei var. hypoglaucum is smaller than that of R. ponticum var. brachycarpum (1C = 0.74 pg) ( Bou Dagher-Kharrat et al., 2013) although both taxa have the same chromosome number 2 n = 2 x = 26 ( Ammal, 1950). This variation in genome size may be due to different internal reference standards: Bou Dagher-Kharrat et al. (2013) used Petunia hybrida (Hook) Vilm. ‘PxPC6’ (1C = 2.85 pg), whereas we utilized Solanum pseudocapsicum (1C = 1.30 pg/1C) ( Temsch et al., 2010). The differences in genome size between the two species might be also due to differential accumulation of the non-coding repetitive DNA, as reported in other plant groups ( Emadzade et al., 2014; Weiss-Schneeweiss et al., 2015; Jang et al., 2018). The genome size variation between species may thus provide an effective taxonomic marker ( Jang et al., 2016; Choi et al., 2019, 2020). In conclusion, this study is the first to report the chromosome number, karyotype, and genome size of the recently reported R. keiskei var. hypoglaucum from a Korean natural population. Although the results in this study show that cytological diversity based on genome size in the examined Korean Rhododendron keiskei var. hypoglaucum species is relatively low, more data incorporating additional species of Rhododendron populations to examine large-scale genomic differences among Korean Rhododendron species are necessary to better understand the genome evolution.
ACKNOWLEDGMENTS
The authors would like to thank Prof. Hanna Weiss-Schneeweiss and Dr. Eva M. Temsch (University of Vienna) for kindly providing the seeds of the reference standard used in genome size estimation. We are also very grateful for Soyun Won at Dadohaehaesang National Park of Korea for their kind help with field survey. This work was supported by grants from the Korean National Park Research Institute, grant number NPRI 2021-09.
Fig. 1.
Photographs of Rhododendron keiskei var. hypoglaucum from Korean natural population. A. Habit. B. Flower. C. Fruits. 
Fig. 2.
Representative chromosomes of Rhododendron keiskei var. hypoglaucum. A, B. Metaphase in mitosis. C. Cut-out karyotypes of the metaphase in mitosis from Fig. 2B. Arrows indicate the satellite (NOR, nucleolar organizer region). 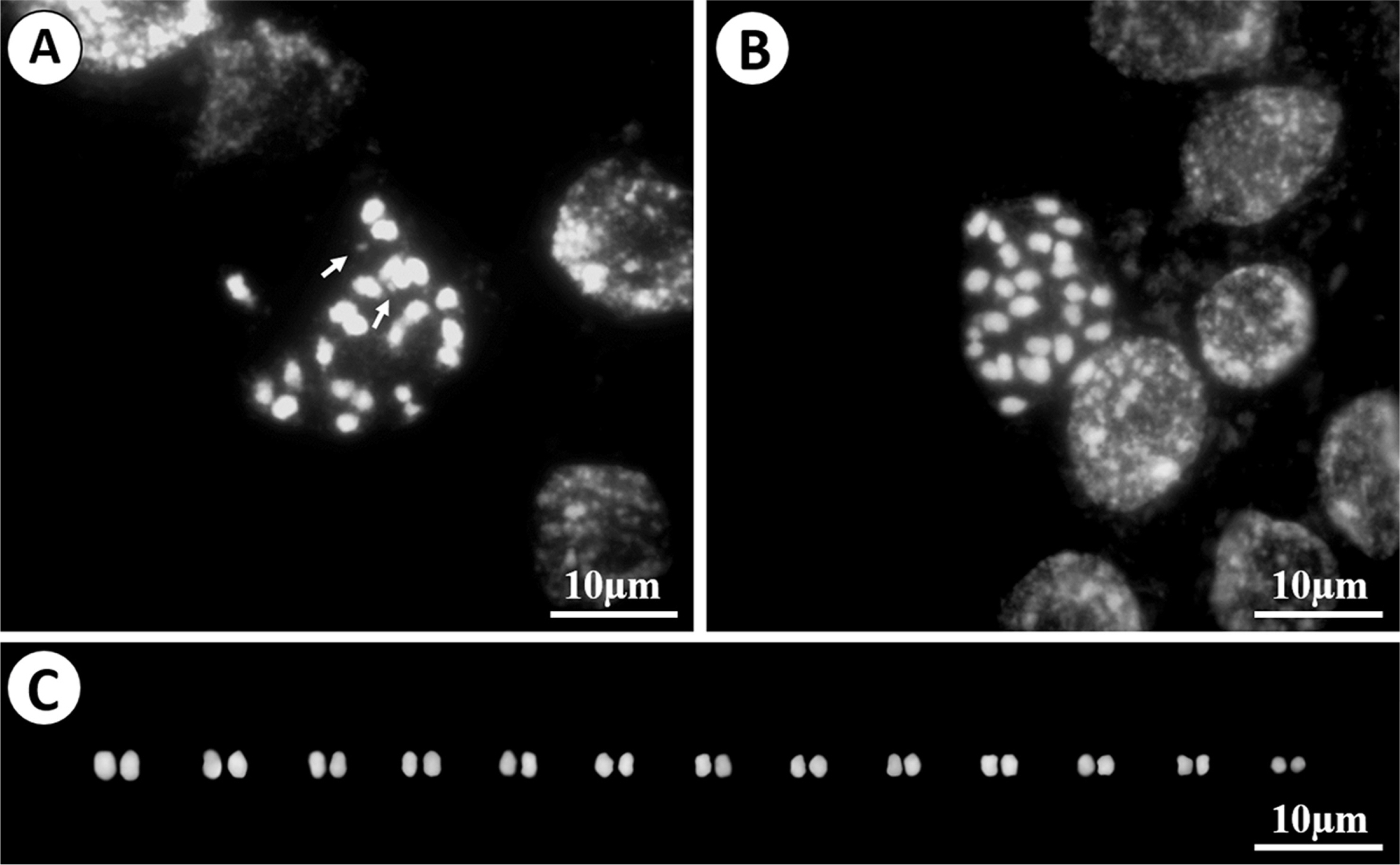
Fig. 3.
Fluorescence histogram (A) and distribution of genome size data (B) of Rhododendron keiskei var. hypoglaucum with internal reference standard used for genome size estimation. 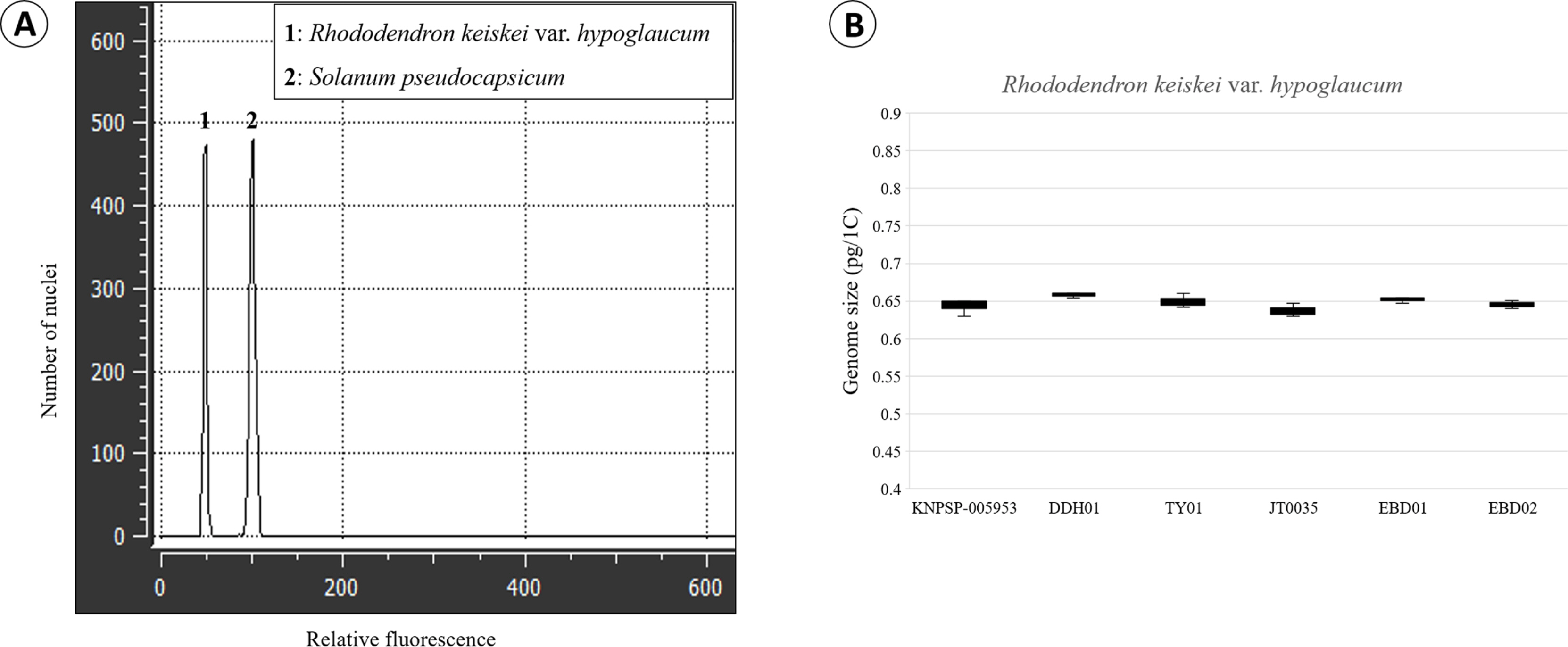
Table 1.
Chromosome number and genome size of Rhododendron keiskei var. hypoglaucum.
|
Accession no. |
2n
|
Genome size (pg) |
|
KNPSP-005953 |
- |
0.6367 ± 0.0094 |
|
DDH01 |
26 |
0.6581 ± 0.0029 |
|
TY01 |
- |
0.6497 ± 0.0076 |
|
JT0035 |
- |
0.6371 ± 0.0074 |
|
EBD01 |
- |
0.6516 ± 0.0030 |
|
EBD02 |
- |
0.6458 ± 0.0053 |
LITERATURE CITED
Ammal, EK. 1950. Polyploidy in the genus Rhododendron
. The Rhododenron Year Book 5: 92-96.
Agudo, AB. Torices, R. Loureiro, J. Castro, S. Castro, M and Álvarez, I. 2019. Genome size variation in a hybridizing diploid species complex in Anacyclus (Asteraceae: Anthemideae). International Journal of Plant Sciences 180: 374-385.  Atkinson, R. Jong, K and Argent, G. 2000. Chromosome numbers of some tropical Rhododendrons (section Vireya). Edinburgh Journal of Botany 57: 1-7.   Bennett, MD. Bhandol, P and Leitch, IJ. 2000. Nuclear DNA amounts in angiosperms and their modern uses–807 new estimates. Annals of Botany 86: 859-909.  Bou Dagher-Kharrat, M. Abdel-Samad, N. Douaihy, B. Bourge, M. Fridlender, A. Siljak-Yakovlev, S and Brown, SC. 2013. Nuclear DNA C-values for biodiversity screening: Case of the Lebanese flora. Plant Biosystems 147: 1228-1237.  Chang, C.-S. 2007. Ericaceae Juss. In The Genera of Vascular Plants of Korea. Park, C. W (ed.), Academy Publishing Co, Seoul. 464-472.
Choi, B. Yang, S. Song, J-H and Jang, T-S. 2019. Karyotype and genome size variation in Ajuga L. (Ajugoideae-Lamiaceae). Nordic Journal of Botany 37: e02337.   Choi, B. Weiss-Schneeweiss, H. Temsch, EM. So, S. Myeong, H-H and Jang, T-S. 2020. Genome size and chromosome number evolution in Korean Iris L. species (Iridaceae Juss.). Plants 9: 1284.    Choi, B. Gang, G-H. Kim, H. Byun, H. Kwak, M. So, S. Myeong, H-H and Jang, T-S. 2021. Cytological study of Cypripedium japonicum Thunb. (Orchidaceae Juss.): An endangered species from Korea. Plants 10: 1978.    Choi, Y. Choi, B and Jang, T-S. 2022. New chromosome counts in Juncus (Juncaceae) taxa from Korea. Cytologia (in press)
Chung, K.-S and Im, H.-T. 2020. Chromosome number report of three Carex sect. Mitratae taxa (Cyperaceae) in Korea. Korean Journal of Plant Taxonomy 50: 361-367.   Chung, K.-S and Chung, GY. 2021. Chromosome numbers of eight Carex taxa in Korea (Cyperaceae). Korean Journal of Plant Taxonomy 51: 192-197.   Emadzade, K. Jang, T-S. Macas, J. Kovařík, A. Novák, P. Parker, J and Weiss-Schneeweiss, H. 2014. Differential amplification of satellite PaB6 in chromosomally hypervariable Prospero autumnale complex (Hyacinthaceae). Annals of Botany 114: 1597-1608.    Fang, MY. Fang, RC. He, MY. Hu, LC. Yang, HP and Chamberlain, DF. 2005. Rhododendron. Flora of China. 14: Wu, Z. Y. Raven, PH (eds.), Science Press, Beijing and Missouri Botanical Garden, St Louis, MO. 260-455.
Greimler, J. Temsch, EM. Xue, Z. Weiss-Schneeweiss, H. Volkova, P. Peintinger, M. Wasowicz, P. Shang, H. Schanzer, I and Chiapella, JO. 2022. Genome size variation in Deschampsia cespitosa sensu lato (Poaceae) in Eurasia. Plant Systematics and Evolution 308: 9.   Gurzenkov, NN. 1973. Studies of chromosome numbers of plants from the south of the Soviet Far East. Komarov Lectures 20: 47-61.
Husband, BC. Baldwin, SJ and Suda, J. 2013. The incidence of polyploidy in natural plant populations: Major patterns and evolutionary processes. Plant Genome Diversity. 2: Physical Structure, Behaviour and Evolution of Plant Genomes. Leitch, IJ. Greilhuber, J. Doležel, J. Wendel, JF (eds.), Springer, Vienna. 255-276.  Ikeda, H. Nam, B-M. Yamamoto, N. Funakoshi, H. Takano, A and Im, H-T. 2021. Chromosome number of myoga ginger ( Zingiber mioga: Zingiberaceae) in Korea. Korean Journal of Plant Taxonomy 51: 100-102.   Jang, T.-S. McCann, J. Parker, JS. Takayama, K. Hong, S.-P. Schneeweiss, GM and Weiss-Schneeweiss, H. 2016. rDNA loci evolution in the genus Glechoma (Lamiaceae). PLoS ONE 11: e0167177.    Jang, T.-S. Parker, JS. Emadzade, K. Temsch, EM. Leitch, AR and Weiss-Schneeweiss, H. 2018. Multiple origins and nested cycles of hybridization result in high tetraploid diversity in the monocot Prospero
. Frontiers in Plant Science 9: 433.    Khan, G. Nolzen, J. Schepker, H and Albach, DC. 2021. Incongruent phylogenies and their implications for the study of diversification, taxonomy, and genome size evolution of Rhododendron
. American Journal of Botany 108: 1957-1981.    Lysak, MA. Berr, A. Pecinka, A. Schmidt, R. McBreen, K and Schubert, I. 2006. Mechanisms of chromosome number reduction in Arabidopsis thaliana and related Brassicaceae species. Proceedings of the National Academy of Sciences of the United States of America 103: 5224-5229.    McCann, J. Jang, T.-S.. Macas, J. Schneeweiss, GM. Matzke, NJ. Novák, P. Stuessy, TF. Villaseñor, JL and Weiss-Schneeweiss, H. 2018. Dating the species network: Allopolyploidy and repetitive DNA evolution in American daisies ( Melampodium sect. Melampodium, Asteraceae). Systematic Biology 67: 1010-1024.    Mitrenina, EY. Erst, A. S.. Peruzzi, L. Skaptsov, MV. Ikeda, H. Nikulin, VY and Wang, W. 2021. Karyotype and genome size variation in white-flowered Eranthis sect. Shibateranthis (Ranunculaceae). PhytoKeys 187: 207-227.   Murin, A. Háberová, I and Žamsran, C. 1984. Further karyological studies of the Mongolian flora. Folia Geobotanica et Phytotaxonomica 19: 29-39.
Otto, SP and Whitton, J. 2000. Polyploid incidence and evolution. Annual Review of Genetics 34: 401-437.   Pessoa, EM. Nollet, F. Magalhães, RF. Viruel, J. Pinheiro, F and Chase, MW. 2022. Nuclear-plastid discordance indicates past introgression in Epidendrum species (Laeliinae: Orchidaceae) with highly variable chromosome numbers. Botanical Journal of the Linnean Society 199: 357-371.   Redpath, LE. Aryal, R. Lynch, N. Spencer, JA. Hulse-Kemp, A. M. Ballington, JR. Green, J. Bassil, N. Hummer, K. Ranney, T and Ashrafi, H. 2022. Nuclear DNA contents and ploidy levels of North American Vaccinium species and interspecific hybrids. Scientia Horticulturae 297: 110955.  Rice, A. Glick, L. Abadi, S. Einhorn, M. Kopelman, NM. Salman-Minkov, A. Mayzel, J. Chay, O and Mayrose, I. 2015. The chromosome counts database (CCDB): A community resource of plant chromosome numbers. New Phytologist 206: 19-26.  Sax, K. 1930. Chromosome stability in the genus Rhododendron
. American Journal of Botany 17: 247-251.   Schubert, I. 2007. Chromosome evolution. Current Opinion in Plant Biology 10: 109-115.   Sliwinska, E. Loureiro, J. Leitch, IJ. Šmarda, P. Bainard, J. Bureš, P. Chumová, Z. Horová, L. Koutecký, P. Lučanová, M. Trávníček, P and Galbraith, DW. 2021. Application-based guidelines for best practices in plant flow cytometry. Cytometry Part A. Advanced online publication.
https://doi.org/10.1002/cyto.a.24499..   Soltis, PS and Soltis, DE. 2009. The role of hybridization in plant speciation. Annual Review of Plant Biology 60: 561-588.   Stebbins, G. L. Jr. 1938. Cytological characteristics associated with the different growth habits in the dicotyledons. American Journal of Botany 25: 189-198.   Stepanov, NV. 1994. Chromosome numbers of some higher plants taxa of the flora of Krasnoyarsk region. Botanicheskii Zhurnal Moscow & Leningrad (St. Petersburg) 79: 135-139.
Temsch, EM. Greilhuber, J and Krisai, R. 2010. Genome size in liverworts. Preslia 82: 63-80.
Temsch, EM. Koutecký, P. Urfus, T. Šmarda, P and Doležel, J. 2021. Reference standards for flow cytometric estimation of abSolute Nuclear Dna Content In Plants. Cytometry Part A. Advanced Online Publication.
Https://doi.org/10.1002/cyto.a.24495
.   Weiss-Schneeweiss, H. Leitch, A. R.. McCann, J. Jang, T.-S and Macas, J. 2015. Employing next generation sequencing to explore the repeat landscape of the plant genome. Next Generation Sequencing in Plant Systematics. Regnum Vegetabile. Hörandl, E. Appelhans, M (eds.), Koeltz Scientific Books, Königstein. 155-179.
Weiss-Schneeweiss, H and Schneeweiss, GM. 2013. Karyotype diversity and evolutionary trends in angiosperms. Plant Genome Diversity. 2: Physical Structure, Behaviour and Evolution of Plant Genomes. Leitch, I. J. Greilhuber, J. Doležel, J. Wendel, JF (eds.), Springer, Vienna. 209-230.  Yang, J.-C.. Kwon, Y-H. Ji, S-J and Shin, C-H. 2015. A new record of Rhododendron keiskei Miq. var. hypoglaucum Sutô & Suzuki (Ericaceae) in Korea. Korean Journal of Plant Taxonomy 45: 239-242.
APPENDICES
Appendix 1.
Previously reported chromosome numbers of Korean Rhododendron species.
|
|














