잡종 기원 녹보리똥나무와 큰보리장나무의 형태학적 및 분자적 다양성 분석 및 평가
적 요:
녹보리똥나무와 큰보리장나무는 형태적 특성에 기반하여 잡종 분류군으로 제안된 바 있으나, 이에 대한 분류학적 실체가 불명확하다. 본 연구에서는 녹보리똥나무와 큰보리장나무의 잡종 기원을 밝히기 위하 여 현장 조사와 표본관에 소장된 표본을 검토하여 형태적 특징을 관찰하였으며, 핵리보솜 구간(internal transcribed spacer, 5S non-transcribed spacer)과 엽록체 구간(matK)의 염기서열을 비교·분석하였다. 형태적 특징을 관찰한 결과, 녹보리똥나무는 보리장나무와, 큰보리똥나무는 통영볼레나무와 형태적으로 유사성을 보였으나, 분자 분석 결과, 녹보리똥나무는 핵리보솜 구간에서 보리장나무와 보리밥나무의 서열의 혼성화가 관찰되었다. 큰보리장나무는 다양한 양상이 관찰되었는데, 일부 개체는 핵리보솜 구간에서 통영볼레나무와 보리밥나무의 서열의 혼성화가, 엽록체 구간에서는 보리밥나무 서열이 관찰되었다. 다른 개체는 핵리보솜 구간 에서는 보리밥나무의 서열을 보였으나, 엽록체 구간에서는 통영볼레나무의 서열이 관찰되어 핵과 엽록체간 불일치를 보였다. 일부 개체는 통영볼레나무와 보리밥나무의 중간형을 보였으나, 핵리보솜과 엽록체 구간 모두 보리밥나무 서열이 관찰되었다. 이러한 결과는 두 종이 잡종기원이며, 부모종 또는 잡종 개체간 교배가 빈번히 일어나는 것을 시사한다.
주요어: 보리수나무속, 엽록체 DNA, 유전적 다양성, 잡종, 핵리보솜 DNA, 형태적 특징
Abstract
The taxonomic identity of Elaeagnus ×maritima and E. ×submacrophylla (Elaeagnaceae) in Korea is unclear, yet they are presumed to be hybrid taxa based on their morphology. To determine their hybrid origins, a morphological analysis (field surveys and specimen examinations) and a molecular analysis involving two nuclear ribosomal DNA (nrDNA) regions (internal transcribed spacer and 5S non-transcribed spacer) and one chloroplast DNA (cpDNA) region (matK) were conducted. The morphological analysis revealed that E. ×maritima showed certain morphological similarities to E. glabra, whereas E. ×submacrophylla showed certain morphological similarities to E. pungens. However, the molecular analysis indicated that E. ×maritima exhibited additive species-specific sites of E. glabra and E. macrophylla in the nrDNA regions. Notably, E. ×submacrophylla showed various aspects, with some individuals exhibiting additive species-specific sites of E. pungens and E. macrophylla in the nrDNA and E. macrophylla sequences in the cpDNA regions, some individuals exhibiting E. macrophylla sequences in the nrDNA and E. pungens sequences in the cpDNA regions, and some individuals displaying E. macrophylla sequences in both the nrDNA and cpDNA regions, despite an intermediate morphology between E. pungens and E. macrophylla. These results indicate that these two species are of hybrid origin and frequently cross between parental and hybrid individuals.
Keywords: chloroplast DNA, Elaeagnus, hybridization, molecular polymorphism, morphological traits, nuclear ribosomal DNA
INTRODUCTION
The genus Elaeagnus L. belongs to the Elaeagnaceae family and comprises approximately 90 species distributed from eastern Asia to Queensland in northeastern Australia, with a few species even occurring in southern Europe and North America ( Qin and Gilbert, 2007). Elaeagnus species are characterized by silvery white or reddish brown peltate scales on most parts of the shoot, alternate leaves, bisexual flowers, 4-lobed calyx, epipetalous, antisepalous, 4-stamens, and 8-riped seed. The genus Elaeagnus has traditionally been divided into two sections: Sempervirentes Serv., which is evergreen, flowering in autumn and fruiting in spring: and Elaeagnus, which is deciduous, flowering in spring and fruiting in autumn ( Servettaz, 1909). Within the section Sempervirentes in Korea, five taxa ( E. glabra Thunb., E. × maritima Koidz., E. macrophylla Thunb., E. × submacrophylla Serv., and E. pungens Thunb.) have been identified. Among these, E. × maritima and E. × submacrophylla have been proposed as hybrid taxa based on their morphological characteristics ( Ohba, 1999; Ko, 2015). Elaeagnus ×maritima was initially considered as a hybrid between E. glabra and E. macrophylla ( Ohba, 1999), whereas Ki (2004) and Koh (2005) regarded it as a synonym of E. glabra based on a comparison of external morphology, trichomes, and pollen. Similarly, E. × submacrophylla is regarded as a hybrid between E. pungens and E. macrophylla ( Servettaz, 1909; Ohba, 1999), but Lee (1996) proposed that it is a hybrid between E. glabra and E. macrophylla instead. On the other hand, while E. × nikaii Nakai, reported on Mt. Mireuksan (Tongyeong, Korea), was regarded as a hybrid between E. pungens and E. macrophylla ( Nakai, 1928), no further research has been conducted on individuals in Mt. Mireuksan. In the taxonomic classification of species within section Sempervirentes in Korea, a consensus among researchers has not yet been reached. This is because of the high morphological variation observed in this group, as well as the possibility of hybridization, which makes species identification challenging. To gain a better understanding of the taxonomic identity of this group of species, it is crucial to genetically examine putative hybrids to determine whether hybridization occurs. Molecular markers such as nuclear ribosomal DNA (nrDNA) regions have proven to be valuable tools for identifying hybrid taxa progenitors because they exhibit biparental inheritance ( Sang et al., 1995; Li, 2006; Du et al., 2009; Les et al., 2009; Hřibová et al., 2011; Kokubugata et al., 2011). The detection of parental genomes in putative hybrid taxa using these markers can provide direct evidence of hybrid speciation. Additionally, the analysis of chloroplast DNA (cpDNA) data can aid in the detection and determination of the direction of hybridization in plants ( Rieseberg et al., 1993; Schwarzbach and Rieseberg, 2002). In this study, the taxonomic identity of putative hybrid taxa was examined by observing their morphological traits, which were compared with the original descriptions, type specimens, and collected and stored specimens. Additionally, two nrDNA regions (internal transcribed spacer [ITS] and 5S non-transcribed spacer [NTS]) and one cpDNA region (matK) were analyzed. Detailed descriptions, illustrations, and photographs were prepared to provide comprehensive documentation of the findings.
MATERIALS AND METHODS
Morphological observation
To compare the morphological characteristics of the putative hybrids and their closely related taxa, we collected specimens from the field sampling in five localities in Korea ( Fig. 1) and deposited them in the herbarium of the Korea National Arboretum (KH). Additional morphological examinations were conducted by observing specimens from the herbariums of the KH, National Institute of Biological Resources (KB), Sungkyunkwan University (SKK), and Yeungnam University (YNUH). Type specimens were obtained by requesting photographs from the herbarium at the University of Tokyo (TI) and the Naturalis Biodiversity Center (L). Morphological traits were observed visually and under a stereomicroscope and measured using a Mitutoyo 500-196-30 Absolute Digimatic Vernier Caliper (Kanagawa, Japan). Additionally, the original descriptions, flora, and monographs were also used as a reference.
DNA extraction and polymerase chain reaction amplification
Leaf material from each individual was collected and dried in silica gel for DNA extraction from voucher specimens (deposited in KH). Total genomic DNA was extracted from the dried leaf material using the DNeasy Plant Mini Kit (QIAGEN, Hilden, Germany) according to the manufacturer’s instructions. The extracted DNA was electrophoresed on a 1% agarose gel to confirm the presence of DNA. The concentration and quality of the DNA were confirmed using a NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA).
Polymerase chain reaction (PCR) amplification of the two nrDNA regions (ITS and 5S NTS) and one cpDNA region ( matK) was performed. The total volume of each PCR mix was 20 μL, comprising 15 μL of distilled water, 1.0 μL of each forward and reverse primer (50 mM), and 1 unit of Taq DNA polymerase master mix (Amplicon, Rødovre, Denmark). The primers used for PCR amplification and the PCR cycle conditions are listed in Table 1. The PCR products were visualized on 1% agarose gels and sequenced using an ABI 3730xl DNA analyzer (Applied Biosystems, Foster City, CA, USA). The determined sequences were deposited in GenBank ( Table 2). The analyzed nucleotide sequences were determined after checking the chromatogram using Geneious R 7.1.9 (Biomatters Ltd., Auckland, New Zealand). All sequences were aligned using MUSCLE software ( Edgar, 2004). Polymorphic sites were identified by overlapping peaks in the chromatogram. For these polymorphic sites, we used the IUPAC ambiguity codes. We used PHASE v.2.1.1. ( Stephens et al., 2001; Stephens and Scheet, 2005) to phase ambiguous sites of nuclear sequences, and format conversion was performed using the SeqPHASE online tool ( Flot, 2010). The phased sequences of the samples were included in the subsequent analyses as sequence version “a” and “b” of the same samples.
Phylogenetic analysis
Maximum likelihood (ML) and Bayesian inference (BI) analyses were conducted for the two nrDNA regions (ITS and 5S NTS) and one cpDNA region ( matK). For the ML analysis, the best-fit model was selected based on the Akaike information criterion implemented in the program jModelTest version 2.1.6 ( Darriba et al., 2012). ML analyses were conducted using IQ-Tree 1.6.12 ( Nguyen et al., 2015), applying a K3P + R2 (in ITS and 5S NTS) and TPM3u + F (in matK) substitution model. 1,000 bootstrap replicates were used to estimate the confidence values of the nodes. The BI analysis used the GTR + I substitution model determined by the Bayesian information criterion ( Schwarz, 1978). The BI inference was performed using MrBayes 3.2.7 ( Ronquist et al., 2012). Four Markov chain Monte Carlo chains were run for 10,000,000 generations each and sampled at every 1,000 generations, starting with a random tree. The convergence of the runs and the estimation of burn-in were checked using Tracer v.1.6 ( Rambaut et al., 2014). Bayesian posterior probabilities (PP) were calculated for the majority consensus tree of all sampled trees after discarding the trees sampled within the burn-in phase (initial 25%) using MrBayes 3.2.7.
RESULTS
Morphological examination
We compared the morphological characteristics of five taxa belonging to the Elaeagnus sect. Sempervirentes. The measured values are listed in Table 3. The prominent morphological traits that were used for identification are habit, twig shape and color, presence or absence of thorns, petiole color and length, leaf shape, color of the abaxial surface of the leaf, shape of the calyx tube, hair type of style, and seed shape ( Fig. 2). Based on the comparison, Elaeagnus × maritima showed similar morphological traits to E. glabra in its scandent stems, reddish brown twigs, abaxial surface of leaves with dense reddish brown scales, gradually constricted calyx tubes, and seeds without beaks. Despite these similarities, E. × maritima showed differences in its angled and rather thick twigs, longer petioles, broader leaves, and 4-angled and broader calyx tubes. E. × submacrophylla showed similar morphological traits to E. pungens in its erect stem, reddish brown twig, abaxial surface of leaves with dense silvery white and sparse brown peltate scales, abruptly constricted calyx tubes, and seeds with beaks. Despite these similarities, E. × submacrophylla showed differences in its leaf shape, calyx shape, and style with white stellate hairs.
nrDNA sequences and phylogenetic analysis
In the ITS region, the sequence length of all examined accessions was 645 bp. Excluding these gaps, the parental species showed sequence variations at eight nucleotide sites. The pairwise-sequence difference between E. glabra and E. macrophylla was five nucleotide substitutions. E. macrophylla and E. pungens differed by four nucleotide substitutions. E. glabra and E. pungens can be distinguished based on five nucleotide substitutions. However, some individuals of E. glabra (JJD178 and JJD180) and E. pungens (TY216), which grew with E. macrophylla, showed polymorphic signals for some species-specific nucleotides, making species identification difficult. Among the putative hybrid species, E. ×maritima exhibited polymorphic ITS signals at those five sites, differentiating itself from E. glabra and E. macrophylla, thus showing sequence additivity. Some individuals of E. ×submacrophylla (GJ378 and TY212) exhibited polymorphic ITS signals at four sites differentiating themselves from E. macrophylla and E. pungens, thus showing sequence additivity. The other individuals of E. ×submacrophylla showed sequences of E. macrophylla ( Table 4). In the 5S NTS, the sequence length for all examined accessions was 200 bp, except for E. glabra which had a length of 199 bp due to a 1 bp deletion at nucleotide site 60. Excluding these gaps, the putative parental species showed sequence variations at 12 nucleotide sites. The pairwise-sequence difference between E. glabra and E. macrophylla was a seven nucleotide substitutions. E. macrophylla and E. pungens differed in 11 nucleotide substitutions. E. glabra and E. pungens can be distinguished based on five nucleotide substitutions. However, similar to the ITS sequence, one individual of E. glabra (JJD178), which grew with E. macrophylla, showed polymorphic signals for some species-specific nucleotides, making species identification difficult. Among the putative hybrid species, E. ×maritima individuals exhibited polymorphic 5S NTS signals at seven sites differentiating itself from E. glabra and E. macrophylla, thus showing sequence additivity. Some individuals of E. ×submacrophylla (GJ378 and TY212) exhibited polymorphic 5S NTS signals at 10 sites differentiating themselves from E. macrophylla and E. pungens, thus showing sequence additivity. Other individuals of E. ×submacrophylla showed squences of E. macrophylla ( Table 4). The combined nrDNA sequences of the five Elaeagnus taxa contained 846 characters, comprising 796 constant, 30 parsimony-uninformative, and 20 parsimony-informative characters. In the combined nrDNA phylogenetic tree, this tree showed that it formed a polytomy. However, these five Elaeagnus taxa were divided into three clades. E. macrophylla, E. ×maritima, and E. ×submacrophylla were formed in a clade (bootstrap support [BS] = 95/PP = 1). E. pungens and E. ×submacrophylla were formed in the second clade (BS = 99/PP = 1). E. glabra and E. ×maritima were formed in the third clade (BS = 92/PP = 1). Each parental taxon was formed in a monophyletic group, and hybrid putative taxa were incorporated into the monophyletic group of each putative parental taxon ( Fig. 3A).
cpDNA sequences and phylogenetic analysis
To trace the plastid donor (maternal origin) of the putative hybrid individuals, the DNA sequences of one plastid marker ( matK) were examined for 24 individuals representing the putative hybrid and its closely related taxa. The size range of matK observed in the examined taxa was 1,015–1,024 bp. In the matK, sequence length for all examined accessions was 1024 bp, except E. pungens and E. ×submacrophylla which had a length of 1,015 bp due to an 8 bp deletion at nucleotide sites 1,395–1,396 and 1,437–1,442. The matK region was useful for identifying the E. pungens and E. macrophylla- E. glabra with 14 nucleotide substitutions, excluding gaps. However, this marker was unable to distinguish between E. glabra and E. macrophylla; therefore, additional markers are needed to clearly distinguish between these two taxa ( Table 5). The aligned matK sequence had a length of 1,525 characters, comprising 1,502 constant, 11 parsimony-uninformative, and 12 parsimony-informative characters. In the matK phylogenetic tree, the putative hybrid individuals and three taxa of the genus Elaeagnus were divided into two clades. E. pungens and E. ×submacrophylla were formed in one clade (BS = 100/PP = 1). E. glabra, E. macrophylla, E. ×maritima, and E. ×submacrophylla were formed in the other clade (BS = 94/PP = 1) ( Fig. 3B).
DISCUSSION
This study confirmed the hybrid origins of E. ×maritima (E. glabra ×E. macrophylla) and E. ×submacrophylla (E. pungens ×E. macrophylla). For E. ×maritima, most individuals exhibited additive species-specific sites of both E. glabra and E. macrophylla in the nrDNA region. However, it was not feasible to determine the plastid donors (maternal origin), because no species-specific sites between E. glabra and E. macrophylla were observed in the cpDNA region.
E. ×submacrophylla exhibits a range of nrDNA and cpDNA characteristics. Some individuals (GJ378 and TY212) displayed sequence additivity and additive species-specific sites for both E. pungens and E. macrophylla in the nrDNA region, and E. macrophylla sequences in the cpDNA region. In contrast, other individuals (TY208 and TY211) showed an E. macrophylla sequence in the nrDNA region, with a polymorphic sequence at some species-specific nucleotides, but possessed E. pungens sequences in the cp region, revealing discrepancies between nrDNA and cpDNA. Furthermore, other individuals (GJ376 and TY213) had sequences of both E. macrophylla in the nrDNA and cpDNA regions, despite their intermediate morphology.
Hybridization can have two main effects on nrDNA sequences, resulting in either additive patterns (both parental ribotypes present) or biased homogenization toward one of the parental lineages. Although divergent ribotypes in hybrid lineages can be quickly homogenized through a process called concerted evolution ( Wendel et al., 1995; Aguilar et al., 1999), some early generations of hybrid lineages can maintain the divergent ribotypes contributed by the parental species ( Siripun and Schilling, 2005; Liu et al., 2009; Cho et al., 2014; Shin et al., 2014; Gil and Kim, 2016). The observed patterns of completely additive polymorphic sites and the lack of new unique mutations in E. × maritima and E. × submacrophylla in relation to their differentiated parental species are consistent with recent or F 1 hybrid formations ( Rieseberg et al., 1993; Nieto Feliner et al., 2004). It is expected that F 2 or higher generations will possess either the nrDNA sequences of one parent only or one predominant nrDNA type, showing incomplete homogenization after a crossing-over event at a single nrDNA locus ( Sang et al., 1995; Wendel et al., 1995). Therefore, intermediate situations between complete additivity and total homogenization can result in the second case. Polyploidization and agamospermy are known to be mechanisms that can retard concerted evolution ( Karvonen and Savolainen, 1993; Suh et al., 1993; Sang et al., 1995; Campbell et al., 1997). However, in Elaeagnus taxa, no cases of polyploidization or agamospermy have been reported. Since Elaeagnus taxa are shrubs, which have relatively longer generation times than herbs and subshrubs, a longer generation time may have contributed to the various aspects observed in the nrDNA region. Three taxa, E. glabra, E. macrophylla, and E. pungens, were morphologically distinct, but there appeared to be incomplete differentiation in their development. This can be a strategic decision for evolutionary success, known as syngameon, a phenomenon observed in Fagaceae species ( Cannon and Petit, 2020; Buck and Flores-Renteria, 2022). In nature, incomplete geographic, ecological (habitat, flowering period), breeding barriers appear to have resulted in frequent hybridization between the parental taxa. First, E. glabra and E. pungens grew in mountainous areas, while E. macrophylla grew near the coast, resulting in a split ecological niche. However, in the mountains near the coast or islands, E. macrophylla grew widely to mountainous areas, and these taxa coexisted. Second, these three taxa share a flowering period from October to November, which increases the likelihood of hybridization or introgression. Finally, hybrid individuals have been observed to produce low fruit and seed yields. Nevertheless, the presence of young hybrid plants indicates that these hybrids can produce fertile seeds. In reproductive biology studies of the genus Elaeagnus, E. umbellate has variability among plants in the separation of male and female floral parts, and selffertilization through autogamy has been observed ( Soley and Sipes, 2021). In order to confirm the breeding system in hybrid taxa, it seems necessary to observe floral morphology, phenology, and pollinators along with pollination experiments. Hybridization and introgression events can complicate the inference of species phylogeny from gene trees, resulting in a polytomy ( Aguilar and Feliner, 2003). Therefore, additional markers and sampling may be necessary to elucidate the relationships among these Elaeagnus species.
Key to five taxa of Elaeagnus section Sempervirentes in Korea
1. Evergreen, flowering in autumn, fruiting in spring; leaves coriaceous (sect. Sempervirentes).
2. Twigs and petioles grayish white or pale brown; abaxial surface midrib of leaves silvery white; style with silvery white peltate scales and white stellate hairs ···················································· E. macrophylla 2. Twigs and petioles reddish brown; midrib of abaxial surface of leaves reddish brown; style without silvery white peltate scales, white stellate hairs present or absent.
3. Stems erect; twigs with thorns (sometimes without thorns in E. ×submacrophylla); abaxial surface of leaves with dense silvery white and sparse brown peltate scales; calyx tube abruptly constricted above the ovary; seed with beak.
4. Leaves elliptic or oblong; calyx tube tubular; style without white stellate hairs ··· E. pungens 4. Leaves ovate-elliptic or broadly elliptic; calyx tube campanulate; style with white stellate hairs ········ ················································· E. ×submacrophylla
3. Stems scandent; twigs without thorns; abaxial surface of leaves with dense reddish brown scales or silvery white peltate scales mixed with reddish brown peltate scales; calyx tube gradually constricted above ovary; seed without beak.
5. Twigs slender, smooth; petioles less than 1 cm long; leaves elliptic or narrowly elliptic; calyx tube slender, not 4-angled ················· E. glabra 5. Twigs rather thick, angular; petioles more than 1 cm long; leaves ovate-elliptic, broadly elliptic, broadly ovate or orbicular; calyx tube rather broad, 4-angled ··························· E. ×maritima
1. Deciduous, flowering in spring, fruiting in autumn; leaves membranous (Sect. Elaeagnus).
Description
Elaeagnus ×maritima Koidz., Bot. Mag. (Tokyo) 31: 133, 1917.—TYPE: JAPAN. Kanagawa Pref., Sagami, Hayama, 3 Nov 1916, K.Hisautsi s.n. (lectotype, TI, seen as photo!, designated here, see Fig. 4A); Musashi, Yokohama, 5 Nov 1916, K.Hisautsi s.n. (syntype, TI, seen as photo!).
Elaeagnus × hisauchii Makino ex Nakai, Bot. Mag. (Tokyo) 32: 223, 1918.—TYPE: KOREA. Jeju-do, 2 Nov 1917, T.Nakai 6352 (syntype, TI, seen as photo!, see Fig. 4B. JAPAN. Musashi, Yokohama, 5 Nov 1916, K.Hisautsi s.n. (syntype, TI, not seen); Yokosuka, J.Matsumura s.n. (syntype, TI, not seen).
Elaeagnus ×liukiuensis Rehder, J. Arnold Arbor. 1: 181, 1920.—TYPE: JAPAN. Okinawa (Liukiu), Okinawa Island, common round Naha, 1 Mar 1917, E.H.Wilson 8159 (holotype, A, seen as photo!; Isotype, K, US, seen as photo!).
Shrubs or vines, evergreen, about 3 m high. Stems scandent; bark shallowly or irregularly longitudinally fissured, gray or grayish brown; lenticels orbicular or elliptic, dark gray; branches gray or grayish brown, glabrous, without thorn; twigs prismatic, brown, grayish brown or gray, 1.5–2.3 mm wide, conspicuous ribs, densely reddish brown peltate scales; winter bud naked, terminal bud oblong or widely ellipsoid, 2.4–3.3 × 1.4–1.9 mm, lateral bud oblong or ovoid, 1.4–2.2 × 0.6–1.4 mm, brown to dark brown, densely reddish brown peltate scales. Leaves alternate, simple; petioles brown, grayish brown or gray, 11.3–19.2 × 1.2–2 mm, densely with reddish brown peltate scales; blades ovate-elliptic, broadly ovate, broadly elliptic or orbicular, 4.8–9.2 × 3.2–5.6 cm, apex acute to acuminate or often caudate with obtuse tip, margin entire, undulate, base obtuse to rounded, coriaceous, adaxial surface dark green, lustrous, sparsely brown peltate scales, glabrescent, abaxial surface yellowish green, densely reddish brown peltate scales or silvery white mixed with reddish brown peltate scales. Inflorescences axillary, fasciculate, 2–5 flowered; pedicels white or pale yellow, 2.6–4.6 × 0.5–0.7 mm, reddish brown peltate scales. Flowers bisexual, incomplete, apetalous, 3.9–6.2 mm in diameter; Calyx tubular, white or pale yellow, densely silvery white peltate scales mixed with reddish brown peltate scales; calyx tube rather broader, gradually constricted above ovary, 4–5.3 × 1.7–3.7 mm, 4-angled; 4-lobed, lobes triangular-ovate, 2– 3.6 × 1.7–3.5 mm, apex acute to acuminate, margin entire; stamens 4, epipetalous, antisepalous, borne in throat of calyx tube; filaments linear, white or yellowish white, ca. 0.4 mm long; anthers basifixed, narrowly oblong to elliptic, yellowish brown or brown, 1.9–2.2 × 0.8–1 mm; ovary epigynous, elliptic, 1.8–2.6 × 1–1.4 mm, 1-loculed; style linear, yellowish green or green, 4.3–5.1 mm long, glabrous or sparsely white stellate hair; stigma, clavate, bended, herkogamy absence or slightly approach hekogamy. Fruits drupe, ellipsoid, red-orange at maturity, 13.7–16.3 × 4.6–6.3 mm, silvery white peltate scales. Seeds oblong, yellowish brown, 13.3–16.1 × 4–5.8 mm, apex gradually constricted, obtuse, surface 8-grooved.
Korean name: Nok-bo-ri-ttong-na-mu (녹보리똥나무).
Phenology: Flowering October to November and fruiting April to May.
Distribution: Japan, Korea (Jeollanam-do, Gyeongsangnam-do, Jeju-do).
Habitat: Streamside, scrublands, and thickets.
Taxonomic notes: Elaeagnus × hisauchii was similar to E. macrophylla by broadly ovate to nearly orbicular leaves and 4-angled campanulate calyx tube. However, it differed from E. macrophylla by reddish brown twigs and petioles, short petioles, small leaves, and gradually constricted calyx tube above the ovary. Therefore, it was previously treated as a synonym of E. × submacrophylla by having the abaxial surface of leaves with dense silvery white and sparse brown peltate scales, style with white stellate hairs ( Nakai, 1928; Lee, 2003). However, E. × hisauchii in Korea was observed not only on Jeju Island, which was previously known as type locality, but also on Heuksan-do Island (see Fig. 4D), and these regions co-occur with E. glabra and E. macrophylla. Also, it differed from E. × submacrophylla by having a vine and gradually constricted calyx tube above the ovary. Consequently, E. × hisauchii in Korea should be considered a synonym of E. ×maritima, which is regarded as a hybrid of E. glabra and E. macrophylla.
Elaeagnus × hisauchii was described by Nakai (1918) based on a specimen from Jeju Island ( T.Nakai 6352, TI; Syntype, see Fig. 4B). However, the specimen has been identified as E. glabra. Rather, another specimen from Jeju Island ( T.Nakai 6351, TI, see Fig. 4C) was identified as E. × hisauchii and best represented the morphology mentioned in the original description. To accurately identify E. × hisauchii, a lectotype among other syntypes should be designated. However, the lectotype could not be designated in this study because other syntypes could not be found.
Additionally, E. ×maritima was assumed as a synonym of E. glabra based on the morphological comparison (external morphology, trichome and pollen) ( Ki, 2004; Koh, 2005), on the other hand, there was a view of integrating into E. ×submacrophylla ( Lee, 1996). However, this species differs from E. glabra by thick and angular twigs, more than 1 cm long petioles, ovate-elliptic, broadly elliptic, broadly ovate or orbicular leaves, and rather broader and 4-angled calyx tube. In addition, it was clearly distinguished from E. × submacrophylla by the abaxial surface of the leaves with dense reddish brown scales or silvery white peltate scales mixed with reddish brown peltate scales, gradually constricted calyx tubes, and seeds without beaks ( Figs. 5, 6).
Representative specimens examined: KOREA. Jeollanam-do: Sinan-gun, Heuksan-myeon, Is. Heuksan, 31 Jan 2008, J.K. Ahn et al., SB20083 (KB); Sinan-gun, Heuksan-myeon, Ye-ri, 22 Oct 2011, B.U. Oh, Sinangun(Daeheuksando)-111022-095–096 (KH); Wando-gun, Bogil-myeon, Is. Bogildo, May 1965, L.M. Kim, KFR20015800–20015802 (KH); Wando-gun, Bogil-myeon, Yesong-ri, Suribong Peak, 7 Apr 2022, B.K. Park et al., Suribong-220407-027–028 (KH); Wando-gun, Gunoe-myeon, 25 Jun 2012, C.J. Oh et al., JN_V2012031 (KH); Wando-gun, Gunoe-myeon, Wando Arboretum, 4 Feb 2005, J.E. Koh & J.H. Kim, B200508052079, s.n. (SKK); Wando-gun, Gunoe-myeon, Daemun-ri, Wando Arboretum, 20 Jul 2013, M.S. Kang et al., 2013JNV027–028 (KH); Wando-gun, Gunoe-myeon, Yeongpung-ri, 3 Oct 2003, K.I. Heo et al. 27983, 28069, 28086, 28087, 28824 (SKK); Wando-gun, Gunoe-myeon, Yeongpung-ri, 23 Oct 2005, J.H. Lee et al. s.n. (SKK); Jindo-gun, Gogun-myeon, Jindo, 1969, L.M. Kim & M.Y. Cho, KFR20015666–20015668 (KH); Jindo-gun, Uisin-myeon, Temp. Ssanggyesa, 19 May 2002, K.I. Heo et al., 26687, 26690, 26691 (SKK); Jindo-gun, Uisinmyeon, Sacheon-ri, Mt. Cheomchalsan, Temp. Ssanggyesa, 6 Nov 2004, K.I. Heo & J.E. Koh s.n. (SKK); Jindo-gun, Uisin-myeon, Sacheon-ri, Unlimsanbang, 4 Nov 2004, K.I. Heo & J.E. Koh s.n. (SKK); Jindo-gun, Jindo-eup, Mt. Cheomchalsan, A path up a mountain of Ariranbi, 5 Nov 2004, K.I. Heo & J.E. Koh s.n. (SKK). Gyeongsangnam-do: Geoje-si, Nambu-myeon, Jeogu-ri, 22 Jun 2010, E.S. Jeon, ESJeon101305 (KH). Jeju-do: Mt. Hallasan, May 1966, L.M. Kim & S.G. Kang, KFR20015663–20015664, s.n. (KH); Jeju-si, Gujwa-eup, Songdang-ri, 21 Sep 2021, B.K. Park et al., Songdangri-210928-001 (KH); Jeju-si, Gujwaeup, Songdang-ri, 17 Oct 2021, K.H. Lee, Songdangri-211017-001–002 (KH); Jeju-si, Gujwa-eup, Songdang-ri, 23 Mar 2022, B.K. Park et al., Songdangri-220323-001–002 (KH); Gujwa-eup, Songdang-ri, 10 Oct 2022, E.S. Kang et al., Songdangri-221012-001–007 (KH); Jeju-si, Arang-dong, Yangji park, 7 Feb 2005, J.H. Kim et al. 054091, 054100, 054102, 054112 (SKK); Jeju-si, Hangyeongmyeon, Cheongsu-ri, Sanyang Keununggot, 2 Nov 2021, B.K. Park et al., Cheongsuri-211102-003–005 (KH); Jeju-si, Haean-dong (Gwangryeongcheon), 28 Sep 2012, D.S. Kim & S.Y. Kim, HALLA1701 (KH); Seogwipo-si, Sanghyo-dong, Donnaeko, 31 Oct 2005, J.H. Kim 04671, 04672, 054252 (SKK); Seogwipo-si, Sanghyo-dong, 23 Mar 2022, B.K. Park et al., Sanghyodong-220323-001–003 (KH); Seogwipo-si, Sanghyo-dong, 26 Apr 2022, Y.J. Jang & K.H. Lee, Sanghyodong-220407-007–008 (KH); Seogwipo-si, Jungmun-dong, Cheonjeyeon Waterfalls, 29 Oct 2010, C.H. Kim et al. 51828 (KB).
Elaeagnus ×submacrophylla Servett., Beih. Bot. Centralbl. 25: 84, 1909.—TYPE: JAPAN. Kyushu , Nagasaki Pref., Nomo-saki, Buerger 385 (lectotype, L, seen as photo!, designated here, see Fig. 7A); Buerger 384 (syntype, L, seen as photo!).
Elaeagnus ×nikaii Nakai, Bot. Mag. (Tokyo) 32: 224, 1918.—TYPE: JAPAN. Yamaguchi Pref., Nagato, Hagi, Tsuruedai, 3 Nov 1917, J. Nikai 2723 (lectotype, TI, seen as photo!, see Fig. 7B; isolecotype, TNS, seen as photo!).
Shrubs, evergreen, about 3 m high. Stems erect; bark shallowly or irregularly longitudinally fissured, gray or grayish brown; lenticels orbicular, dark gray; branches dark gray or grayish brown, glabrous, thorn present or absent; twigs terete, brown, grayish brown or gray, 1.6–2.2 mm wide, densely reddish brown peltate scales; winter bud naked, brown to dark brown, densely reddish brown peltate scales, terminal bud oblong or widely ellipsoid, lateral bud oblong or ovoid, 1.5–2.3 × 0.7–1 mm. Leaves alternate, simple; petioles brown, grayish brown or gray, 10.6–19.1 × 1.1–2.7 mm, densely reddish brown peltate scales; blades ovate-elliptic to broadly elliptic, 5.6–9.3 × 3.7–5.1 cm, apex acute to acuminate or often caudate with obtuse tip, margin entire, undulate, base obtuse to rounded, coriaceous, adaxial surface dark green, lustrous, sparsely silvery white peltate scales, glabrescent, reticulate vein inconspicuous when they are dry, abaxial surface yellowish green, densely silvery white peltate scales, punctured with reddish brown peltate scales. Inflorescences axillary, fasciculate, 1–3 flowered; pedicels white or pale yellow, 2.4–3.9 × 0.6–0.7 mm, reddish brown peltate scales. Flowers bisexual, incomplete, apetalous, 5.5–9.5 mm in diameter; Calyx campanulate, white or pale yellow, silvery white peltate scales and reddish brown peltate scales; calyx tube abruptly constricted above ovary, 4.9–6.2 × 2.7–4.2 mm, 4-angled; 4-lobed, lobes triangular-ovate, 2.7–4.7 × 2.2–4.4 mm, apex acute to acuminate, margin entire; stamens 4, epipetalous, antisepalous, borne in throat of calyx tube; filaments linear, white or yellowish white, 0.4–0.5 mm long; anthers basifixed, narrowly oblong to elliptic, yellowish brown or brown, 2.9–3.2 × 1–1.3 mm; ovary epigynous, elliptic, 2.2– 3.1 × 1.3–1.8 mm, 1-loculed; style linear, yellowish green or green, 8–9.3 mm long, whitish stellate hairs; stigma beyond stamens, clavate, bended, herkogamy absence or slightly approach hekogamy. Fruits drupe, ellipsoid, red-orange at maturity, 11.7–12.4 × 5.2–5.8 mm, silvery white peltate scales. Seeds oblong, yellowish brown, 10.7–11.2 × 4.2–4.8 mm, apex attenuate, beaked, surface 8-grooved.
Korean name: Keun-bo-ri-jang-na-mu (큰보리장나무).
Phenology: Flowering October to November and fruiting April to May.
Distribution: Japan, Korea (Gyeongsangnam-do).
Habitat: Streamside, scrublands, and thickets.
Taxonomic notes: Elaeagnus ×submacrophylla was described by Servettaz (1909) and is assumed to be a hybrid of E. pungens and E. macrophylla. However it was recognized as a hybrid of E. glabra and E. macrophylla by Nakai (1928), who treated E. × hisauchii Nakai as a synonym of E. ×submacrophylla, and the hybrid of E. pungens and E. macrophylla was treated as E. × nikaii Nakai ( Lee, 1996; Lee, 2003; Ohashi, 2003). In a recent taxonomic treatment, E. × hisauchii was treated as a synonym of E. ×maritima, which is regarded as a hybrid of E. glabra and E. macrophylla, and E. × nikaii was treated as a synonym of E. ×submacrophylla, which is regarded as a hybrid of E. pungens and E. macrophylla ( Ohashi, 2003; Ko, 2015). Elaeagnus ×submacrophylla in Japan has been known to be distributed in regions where E. pungens and E. macrophylla co-occur ( Ohashi and Yoshida, 2003). But E. ×submacrophylla previously reported in Korea is confused about the distribution in that it was a region where E. glabra and E. macrophylla co-occur or only E. macrophylla occurs ( Ko, 2015). As a result of the field survey and reclassification of the specimens, those identified as E. ×submacrophylla in Korea were confirmed to be either E. macrophylla or E. × hisauchii (= E. ×maritima). In addition, E. × nikaii in Korea was not only previously reported on Mt. Mireuksan, Tongyeong where E. pungens and E. macrophylla co-occur, but also was morphologically similar to E. macrophylla, but differed by reddish brown twigs, petioles, midrib of abaxial surface of leaves, style with no silvery white peltate scales and only white stellate hairs ( Figs. 8, 9). Therefore, E. × nikaii previously reported in Korea is considered to be E. ×submacrophylla.
Representative specimens examined: KOREA. Gyeongsangnam-do: Geoje-si, Dundeok-myeon, Sanbangri, Temp. Bohyeonsa, 30 Mar 2022, B.K. Park et al., Bohyeonsa-220330-001–002 (KH); Dundeok-myeon, Sanbang-ri, Temp. Bohyeonsa, 19 Oct 2022, Y.J. Jang et al., Bohyeonsa-221019-003–004 (KH); Tongyeong-si, Bongpyeong-dong, Mt. Mireuksan, 26 Apr 2005, J.H. Lee et al. 044754–044756, 046451–046453, 054328, s.n. (SKK); Tongyeong-si, Bongpyeong-dong, Mt. Mireuksan, 27 Oct 2021, B.K. Park et al., Mireuksan-211027-001–002 (KH); Tongyeong-si, Bongpyeong-dong, Mt. Mireuksan, 31 Mar 2022, B.K. Park et al., Mireuksan-220331-001–005 (KH); Tongyeong-si, Bongpyeong-dong, Mt. Mireuksan, 20 Oct 2022, B.K. Park et al., Mireuksan-220331-001–005 (KH); Tongyeong-si, Bongpyeong-dong, Mt. Mireuksan, 20 Oct 2022, Y.J. Jang et al., Mireuksan-220331-004 (KH).
ACKNOWLEDGMENTS
We are grateful to the persons concerned at the KB, SKK, and YNUH herbaria for permitting the examination of specimens, Dr. Akiko Shimizu (TI Herbarium) and Roxali Bijmoer (Naturalis Biodiversity Center) for providing images and allowing us to use, Hyeryun Jo for preparing the line drawings. This study was supported by the Korea National Arboretum (KNA1-1-18, 15-3).
Fig. 1.
Investigated localities (TY, Tongyeong; GJ, Geoje; JJD, Jejudo Island; WD, Wando; HSD, Heuksando Island). 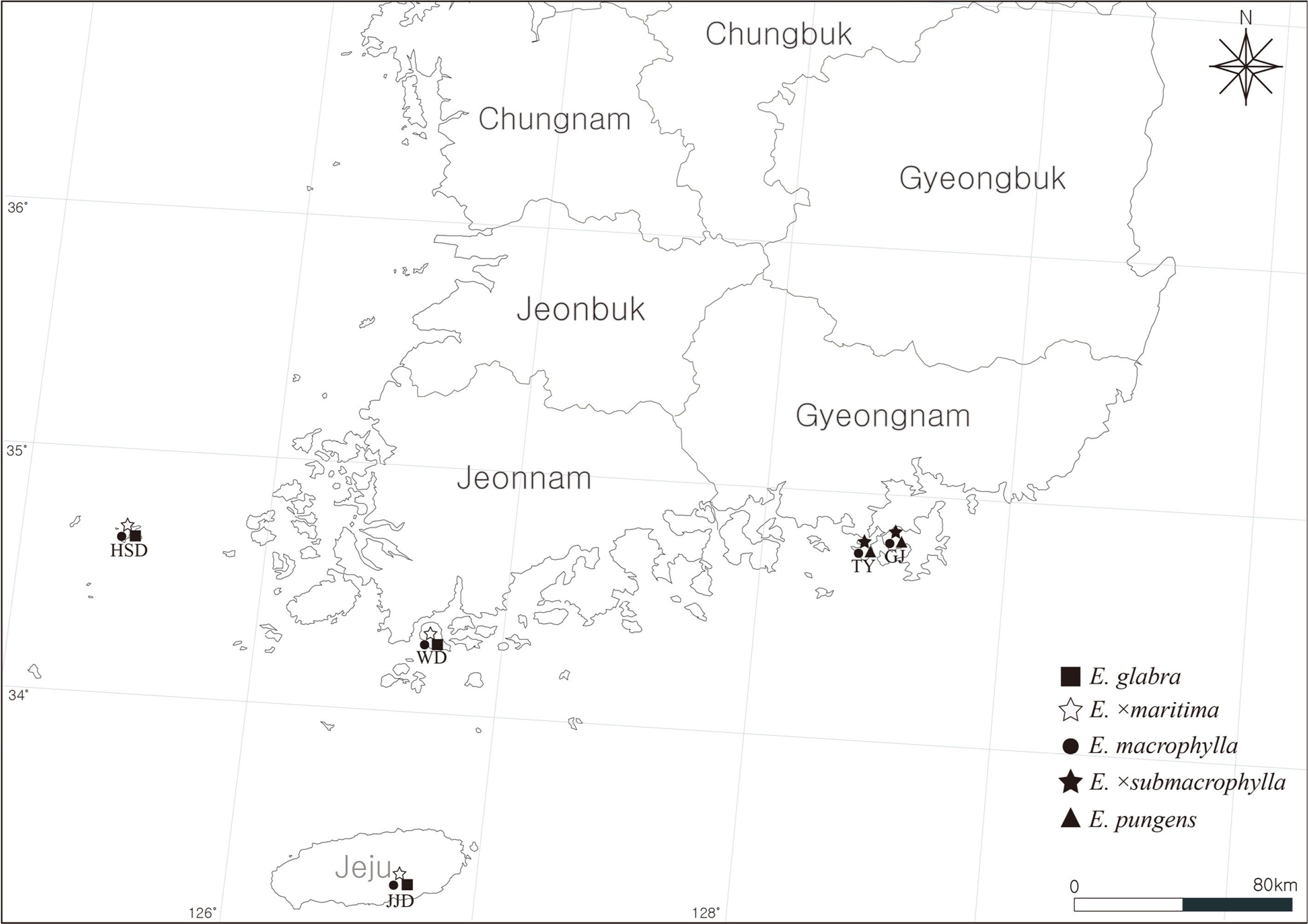
Fig. 2.
Illustrations of the major morphological characters among Elaeagnus section Sempervirentes taxa in Korea. A. Twig. B. Leaf (abaxial). C. Flower (longitudinally section). D. Seed. Illustrations by Hyeryun Jo. 
Fig. 3.
Bayesian tree of Elaeagnus sect. Sempervirentes in Korea, (A) based on nrDNA (ITS + 5S NTS) and (B) cpDNA (matK). Numbers above branches represent bootstrap support and posterior probability (ML/BS), respectively. Putative hybrid species are indicated by black and white star shapes. nrDNA, nuclear ribosomal DNA; ITS, internal transcriped spacer; NTS, nontranscribed spacer; cpDNA, chloroplast DNA; ML, maximum likelihood; BS, bootstrap support. 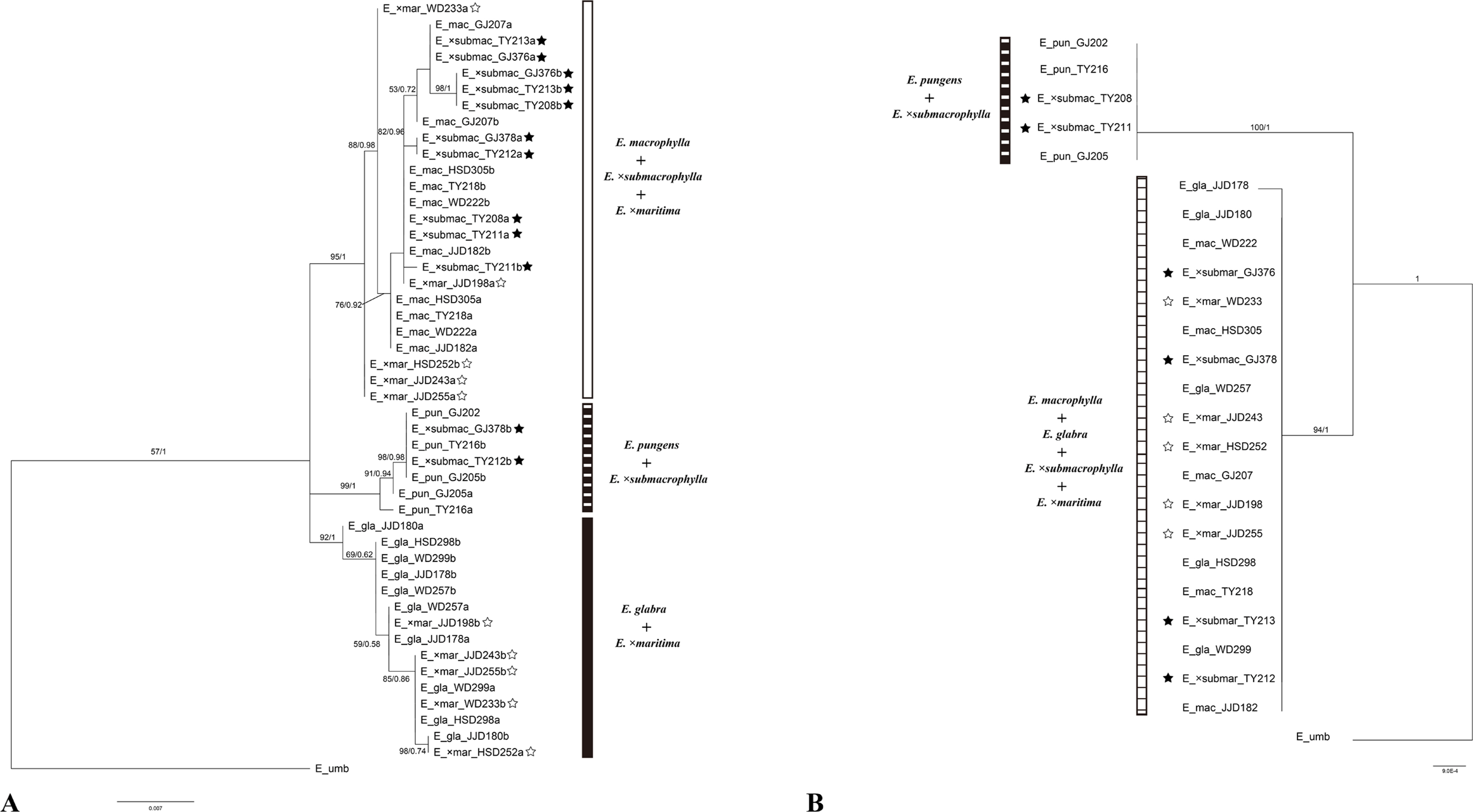
Fig. 4.
Specimens of Elaeagnus ×maritima Koidz. and E. ×hisauchii Makino ex Nakai. A. Lectotype of E. ×maritima (K.Hisauchi s.n., TI, designated here). B. Syntype of E. ×hisauchii (T.Nakai 6352, TI) identified as E. glabra. C. E. ×hisauchii (Jeju-do, Korea, 3 Nov 1917, T.Nakai 6351, TI). D. E. ×maritima (Is. Heuksando, Sinan-gun, Jeollanam-do, Korea, 22 Oct 2011. B.U.Oh 111022-096, KH) previously identified as E. ×submacrophylla. 
Fig. 5.
Photographs of Elaeagnus ×maritima Koidz. A. Habit. B. Bark. C. Branch. D. Twig. E. Leaf (adaxial). F. Leaf (abaxial). G. Inflorescence. H. Flower (lateral view). I. Flower (top view). J. Pistil. K. Fruit. L. Seed. 
Fig. 6.
Illustrations of Elaeagnus ×maritima Koidz. A. Flowering habit. B. Fruiting habit. C. Twig. D. Winter bud. E. Leaf (adaxial). F. Leaf (abaxial). G. Leaf margin (adaxial). H. Leaf margin (abaxial). I. Petiole (adaxial). J. Leaf midrib (abaxial). K–M. Peltate scales. N. Flower (lateral view). O. Flower longitudinally section. P. Flower (top view). Q. Pistil. R. Stamen (adaxial). S. Stamen (abaxial). T. Fruit. U. Endocarp. V. Endocarp transverse section W. Seed. Illustrations by Hyeryun Jo. 
Fig. 7.
Specimens of Elaeagnus ×submacrophylla Servett. and E. ×nikaii Nakai. A. Lectotype of E. ×submacrophylla (Buerger 385, L, designated here). B. Lectotype of E. ×nikaii (J.Nikai 2723, TI).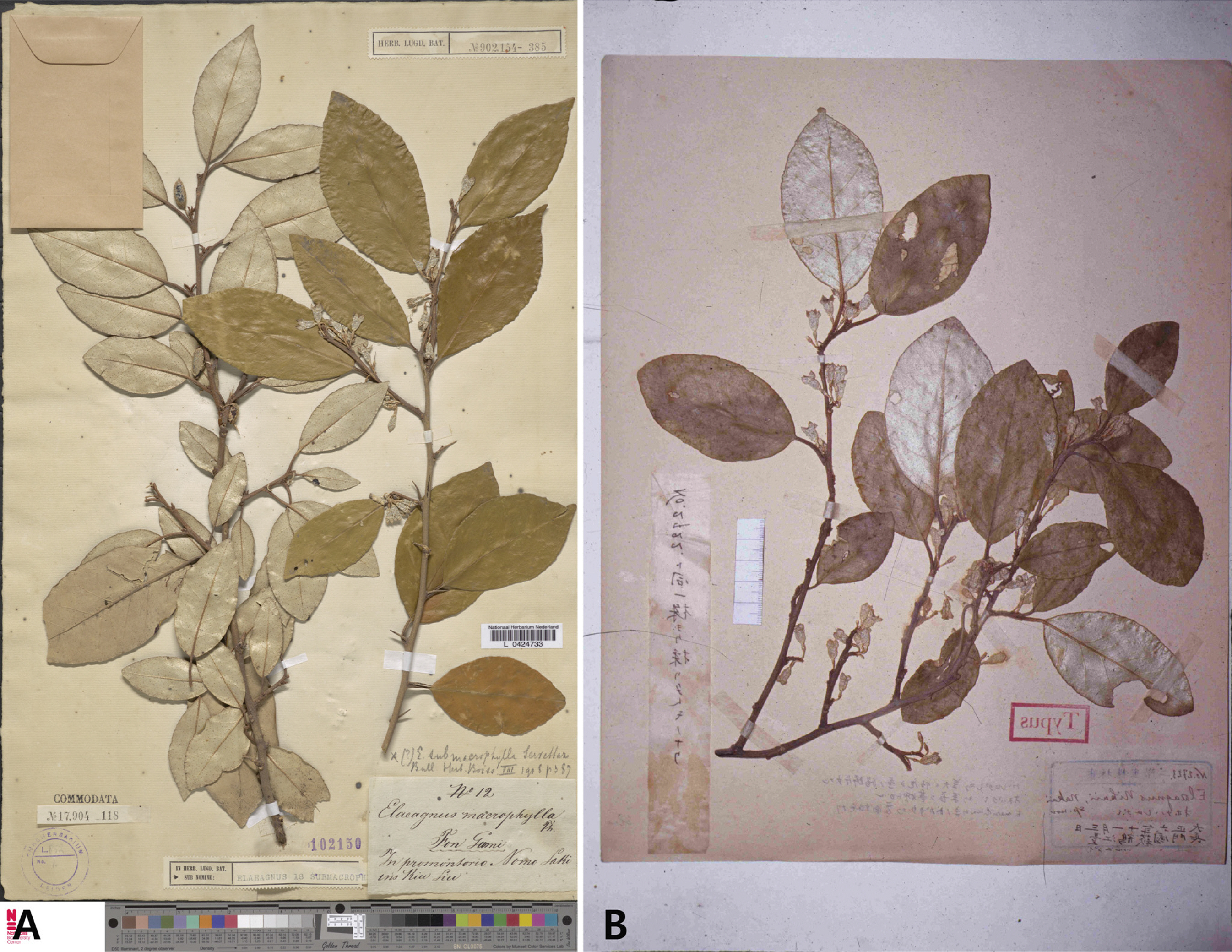 
Fig. 8.
Photographs of Elaeagnus ×submacrophylla Servett. A. Habit. B. Bark. C. Branch. D. Twig. E. Leaf (adaxial). F. Leaf (abaxial). G. Inflorescence. H. Flower (lateral view). I. Flower (top view). J. Pistil. K. Fruit. L. Seed. 
Fig. 9.
Illustrations of Elaeagnus ×submacrophylla Servett. A. Flowering habit. B. Fruiting habit. C. Twig (with thorn). D. Leaf (adaxial). E. Leaf (abaxial). F. Leaf margin (adaxial). G. Leaf margin (abaxial). H. Petiole (adaxial). I. Leaf midrib (abaxial). J–L. Peltate scales. M. Flower (lateral view). N. Flower longitudinally section. O. Flower (top view). P. Pistil. Q. Stamen (adaxial). R. Stamen (abaxial). S. Fruit. T. Endocarp. U. Endocarp transverse section. V. Seed. Illustrations by Hyeryun Jo. 
Table 1.
List of primers and PCR conditions used in the present study.
|
Region |
Primer name |
Primer sequence (5′ – 3′) |
Reference |
PCR condition |
|
ITS |
ITS 5 |
GGA AGT AAA AGT CGT AAC AAG G |
White et al. (1990)
|
95°C for 5 min, 35 cycles of 94°C for 20 s, 56°C for 20 s, 72°C for 1 min, 72°C for 5 min |
|
ITS 4 |
TCC TCC GCT TAT TGA TAT GC |
|
5S NTS |
5S1 |
GGA TGG TGA CCT CCC GGG AAG TCC |
Falistocco et al. (2007)
|
95°C for 5 min, 35 cycles of 95°C for 20 s, 60°C for 20 s, 72°C for 30 s, 72°C for 5 min |
|
5S2 |
CGC TTA ACT GCG GAG TTC TGA TGG G |
|
matK |
trnK685F |
GTA TCG CAC TAT GTA TCA TTT GA |
Wojciechowski et al. (2004)
|
95°C for 5 min, 35 cycles of 94°C for 20 s, 55°C for 20 s, 72°C for 1 min, 72°C for 5 min |
|
matK832R |
TTG CAT AGA AAT AGA TTC GCT CAA A |
|
matK4La |
CCT TCG ATA CTG GGT GAA AGA T |
|
matK1932Ra |
CCA GAC CGG CTT ACT AAT GGG |
|
matK1100L |
TTC AGT GGT ACG GAG TCA AAT G |
|
trnK2R |
CCC GGA ACT AGT CGG ATG G |
Table 2.
List of taxa used for nrITS and cpDNA sequences analysis with voucher and GenBank accession numbers.
|
Taxon |
Locality and voucher specimen |
nrDNA |
cpDNA |
|
|
ITS |
5S NTS |
matK |
|
Elaeagnus glabra
|
Myosanbong Peak, Gujwa-eup, Jeju-si, Jeju-do, Korea Myosanbong-220323-001 (JJD178) |
OQ718935 |
OQ714343 |
OQ714367 |
|
Sanghyo-dong, Seogwipo-si, Jeju-do, Korea Sanghyodong-220323-007 (JJD180) |
OQ718936 |
OQ714344 |
OQ714368 |
|
Wando Arboretum, Daemun-ri, Wando-gun, Jeollanam-do, Korea 2013JNV027 (WD257) |
OQ718937 |
OQ714345 |
OQ714379 |
|
Wando-eup, Wando-gun, Jeollanam-do, Korea S-0568 (WD299) |
OQ718939 |
OQ714347 |
OQ714381 |
|
Ye-ri, Heuksan-myeon, Sinan-gun, Jeollanam-do, Korea Sinangun(Daeheuksando)-171022-111 (HSD298) |
OQ718938 |
OQ714346 |
OQ714380 |
|
E. macrophylla
|
Bomok, Bomok-dong, Seogwipo-si, Jeju-do, Korea Bomok-220323-001 (JJD182) |
OQ718944 |
OQ714352 |
OQ714369 |
|
Temp. Bohyeonsa, Sanbang-ri, Geoje-si, Gyeongsangnam-do, Korea Bohyeonsa-220330-001 (GJ207) |
OQ718949 |
OQ718949 |
OQ714370 |
|
Temp. Milaesa, Tongyeong-si, Gyeongsangnam-do, Korea Milaesa-220331-001 (TY218) |
OQ718945 |
OQ714353 |
OQ714371 |
|
Gugyedeung, Wando-gun, Jeollanam-do, Korea Gugyedeung-220406-001 (WD222) |
OQ718946 |
OQ714354 |
OQ714372 |
|
Ye-ri, Heuksan-myeon, Sinan-gun, Jeollanam-do, Korea Heuksando(Yeri)-171009-012 (HSD305) |
OQ718947 |
OQ714355 |
OQ714382 |
|
E. pungens
|
Hogok437, Suryeok-ri, Geoje-si, Gyeongsangnam-do, Korea Suryeokri-221019-001 (GJ202) |
OQ718950 |
OQ718950 |
OQ714386 |
|
Dundeok Fortress, Georim-ri, Geoje-si, Gyeongsangnam-do, Korea Dundeok Fortress-220330-001 (GJ205) |
OQ718951 |
OQ718951 |
OQ714387 |
|
Mt. Mireuksan, Bongpyeong-dong, Tongyeong-si, Gyeongsangnam-do, Korea Mireuksan-220331-011 (TY216) |
OQ718952 |
OQ714360 |
OQ714388 |
|
E. ×maritima
|
Songdang-ri, Gujwa-eup, Jeju-si, Jeju-do, Korea Songdangri-220323-001 (JJD198) |
OQ718948 |
OQ714356 |
OQ714373 |
|
Sanghyo-dong, Seogwipo-si, Jeju-do, Korea Sanghyodong-220426-007 (JJD243) |
OQ718941 |
OQ714349 |
OQ714374 |
|
Gwangnyeongcheon River, Haean-dong, Jeju-si, Jeju-do, Korea HALLA1701 (JJD255) |
OQ718942 |
OQ714350 |
OQ714375 |
|
Suribong Peak, Bogil-myeon, Wando-gun, Jeollanam-do, Korea Suribong-220407-027 (WD233) |
OQ718940 |
OQ714348 |
OQ714383 |
|
Ye-ri, Heuksan-myeon, Sinan-gun, Jeollanam-do, Korea Sinangun(Heuksando)-111022-095 (HSD252) |
OQ718943 |
OQ714351 |
OQ714385 |
|
E. ×submacrophaylla
|
Temp. Bohyeonsa, Sanbang-ri, Geoje-si, Gyeongsangnam-do, Korea Bohyeonsa-221019-003 (GJ376) |
OQ718955 |
OQ714363 |
OQ714384 |
|
Temp. Bohyeonsa, Sanbang-ri, Geoje-si, Gyeongsangnam-do, Korea Bohyeonsa-221019-004 (GJ378) |
OQ718958 |
OQ714366 |
OQ714376 |
|
Mt. Mireuksan, Bongpyeong-dong, Tongyeong-si, Gyeongsangnam-do, Korea Mireuksan-220331-001 (TY208) |
OQ718953 |
OQ714361 |
OQ714389 |
|
Mt. Mireuksan, Bongpyeong-dong, Tongyeong-si, Gyeongsangnam-do, Korea Mireuksan-220331-002 (TY211) |
OQ718956 |
OQ714364 |
OQ714390 |
|
Mt. Mireuksan, Bongpyeong-dong, Tongyeong-si, Gyeongsangnam-do, Korea Mireuksan-220331-003 (TY212) |
OQ718957 |
OQ714365 |
OQ714377 |
|
Mt. Mireuksan, Bongpyeong-dong, Tongyeong-si, Gyeongsangnam-do, Korea Mireuksan-220331-004 (TY213) |
OQ718954 |
OQ714362 |
OQ714378 |
Table 3.
Comparison of the morphological characters among Elaeagnus section Sempervirentes taxa in Korea.
|
E. glabra
|
E. ×maritima
|
E. macrophylla
|
E. ×submacrophylla
|
E. pungens
|
|
Stem |
Scandent |
Scandent |
Scandent |
Erect |
Erect |
|
Thorn |
Absent |
Absent |
Absent |
Present or absent |
Present |
|
Twig |
Slender, smooth, reddish brown |
Rather thick, angular, reddish brown |
Rather thick, angular, grayish white or pale brown |
Rather thick, angular, reddish brown |
Slender, smooth, reddish brown |
|
Petiole |
Reddish brown |
Reddish brown |
Grayish white or pale brown |
Reddish brown |
Reddish brown |
|
Length of petiole (cm) |
0.7–1.3 |
1.1–1.7 |
1.2–2.3 |
0.9–1.5 |
0.7–1.4 |
|
Leaf shape |
Narrowly elliptic to elliptic |
Ovate-elliptic, elliptic, broadly elliptic or orbicular |
Broadly ovate, oval or nearly orbicular |
Ovate-elliptic to broadly elliptic |
Elliptic or oblong |
|
Leaf size (cm) |
3.8–8.2 × 1.8–4.3 |
4.2–9.2 × 3.2–5.6 |
5–10.8 × 2.9–8.5 |
5.8–13.9 × 3.2–9 |
4.3–9.2 × 1.7–4.1 |
|
Abaxial surface of leaf |
Densely reddish brown peltate scales or reddish brown mixed with silvery white peltate scales |
Densely reddish brown peltate scales or silvery white mixed with reddish brown peltate scales |
Densely silvery white peltate scales, mixed with brown peltate scales, glabrescent |
Silvery white peltate scales, punctured with reddish brown peltate scales |
Densely silvery white peltate scales, punctured with reddish brown peltate scales. |
|
Calyx shape |
Funnelform-tubular, gradually constricted |
Tubular, gradually constricted |
Campanulate, abruptly constricted |
Campanulate, abruptly constricted |
Tubular, abruptly constricted |
|
Calyx size (mm) |
4.1–6.1 × 1.5–2.4 |
3.7–5.6 × 2–3.4 |
4.8–6.4 × 3.4–3.8 |
5.5–6.2 × 3.5–4.1 |
4.8–8.6 × 2.1–4.1 |
|
Style |
Glabrous or rarely white stellate hairs |
Glabrous or white stellate hairs |
Silvery white peltate scales and white stellate hairs |
White stellate hairs |
Glabrous |
|
Achene |
Apex gradually constricted, obtuse |
Apex gradually constricted, obtuse |
Apex gradually constricted, obtuse |
Apex attenuate, beaked |
Apex attenuate, beaked or obtuse |
Table 4.
Nuclear species-specific sites in Elaeagnus section Sempervirentes taxa in Korea.
|
Taxon |
Voucher |
Locality |
ITS |
5S NTS |
|
|
|
112 |
125 |
192 |
466 504 |
532 |
630 |
631 |
60 |
62 |
66 |
72 |
76 |
87 |
88 |
111 |
114 |
144 |
157 |
191 |
|
E. glabra
|
JJD178 |
Jeju |
G/C |
T |
T/C |
T |
T |
C |
A |
T/G |
- |
A |
A |
C |
A/C |
A/G |
T |
T/C |
C |
C/G |
A/C |
A |
|
JJD180 |
Jeju |
A/C |
T/C |
T |
T/C |
T |
C |
A |
G |
- |
A |
A |
C |
C |
A |
T |
T |
C |
C |
A/C |
A |
|
WD257 |
Wando |
G |
T |
T |
T |
T |
C |
A |
G |
- |
A |
A |
C |
C |
A |
T |
T |
C |
C |
A/C |
A |
|
WD299 |
Wando |
A/G |
T |
T/C |
T |
T |
C |
A |
G |
- |
A |
A |
C |
C |
A |
T |
T |
C |
C |
A/C |
A |
|
HSD298 |
Huksando |
A/G |
T |
T/C |
T |
T |
C |
A |
G |
- |
A |
A |
C |
C |
A |
T |
T |
C |
C |
A/C |
A |
|
E. macrophylla
|
JJD182 |
Jeju |
C |
C |
C |
C |
T |
C |
A/C |
T/G |
A/T |
C |
A |
C |
A |
G |
T |
C |
C |
G |
C |
G/A |
|
TY218 |
Tongyeong |
C |
C |
C |
C |
T |
C |
A/C |
T/G |
A/T |
C |
A |
C |
A |
G |
T |
C |
C |
G |
C |
G/A |
|
WD222 |
Wando |
C |
C |
C |
C |
T |
C |
A/C |
T/G |
A/T |
C |
A |
C |
A |
G |
T |
C |
C |
G |
C |
G/A |
|
HSD305 |
Huksando |
C |
C |
C |
C |
T |
C |
A/C |
T/G |
A/T |
C |
A |
C |
A |
G |
T |
C |
C |
G |
C |
G/A |
|
E. pungens
|
GJ202 |
Geoje |
C |
C |
C |
C |
C |
A |
C |
G |
T |
A |
T |
G |
C |
A |
C/T |
T |
G |
C |
C |
A |
|
GJ205 |
Geoje |
C |
C |
C |
C |
C |
A/C |
C |
G |
T |
A |
T |
G |
C |
A |
C/T |
T |
G |
C |
C |
A |
|
TY216 |
Tongyeong |
C |
C |
C |
C |
C |
A/C |
C |
T/G |
T |
A |
A/T |
G/C |
C |
A |
C/T |
T |
G |
C |
C |
A |
|
E. ×maritima
|
JJD198 |
Jeju |
G/C |
T/C |
T/C |
T/C |
T |
C |
A |
T/G |
A/T |
A/C |
A |
C |
A/C |
A/G |
T |
T/C |
C |
C/G |
A/C |
G/A |
|
JJD243 |
Jeju |
G/C |
T/C |
T/C |
T/C |
T |
C |
A/C |
T/G |
- |
A/C |
A |
C |
A/C |
A/G |
T |
T/C |
C |
C/G |
A/C |
A |
|
JJD255 |
Jeju |
G/C |
T/C |
T/C |
T/C |
T |
C |
A |
T/G |
- |
A/C |
A |
C |
A/C |
A/G |
T |
T/C |
C |
C/G |
A/C |
A |
|
WD233 |
Wando |
C |
T/C |
T/C |
T/C |
T |
C |
A/C |
T/G |
- |
A/C |
A |
C |
A/C |
A/G |
T |
T/C |
C |
C/G |
A/C |
A |
|
HSD252 |
Huksando |
A/C |
T/C |
C |
T/C |
T |
C |
A/C |
G |
- |
A/C |
A |
C |
A/C |
A/G |
T |
T/C |
C |
C/G |
A/C |
A |
|
E. ×submacrophaylla
|
GJ376 |
Geoje |
C |
C |
C |
C |
T/C |
A/C |
C |
G |
T |
C |
A |
C |
A |
G |
T |
C |
C |
G |
C |
G/A |
|
GJ378 |
Geoje |
C |
C |
C |
C |
T/C |
A/C |
A/C |
T/G |
T |
A/C |
A/T |
G/C |
A/C |
G/A |
C/T |
T/C |
C/G |
G/C |
C |
G/A |
|
TY208 |
Tongyeong |
C |
C |
C |
C |
T/C |
A/C |
A/C |
T/G |
T |
a/C |
A |
C |
A |
G |
T |
C |
C |
G |
C |
G/A |
|
TY211 |
Tongyeong |
C |
C |
C |
C |
T |
C |
A/C |
T/G |
T |
C |
A |
C |
A |
G |
T |
C |
C |
G |
C |
G/A |
|
TY212 |
Tongyeong |
C |
C |
C |
C |
T/C |
A/C |
A/C |
T/G |
T |
A/C |
A/T |
G/C |
A/C |
G/A |
C/T |
T/C |
C/G |
G/C |
C |
G/A |
|
TY213 |
Tongyeong |
C |
C |
C |
C |
T/C |
A/C |
C |
G |
T |
C |
A |
C |
A |
G |
T |
C |
C |
G |
C |
G/A |
Table 5.
Plastid species-specific sites in Elaeagnus section Sempervirentes taxa in Korea.
|
Taxon |
Voucher |
Locality |
matK |
|
|
40 |
73 |
75 |
156 |
237 |
704 |
790 |
1020 |
1022 |
1033 |
1074 |
1230 |
1395–1397 |
1437–1442 |
1489 |
1507 |
|
E. glabra
|
SOK-178 |
Jeju |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
SOK-180 |
Jeju |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
SOK-257 |
Wando |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
SOK-299 |
Wando |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
SOK-298 |
Huksando |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
E. macrophylla
|
SOK-182 |
Jeju |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
SOK-218 |
Tongyeong |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
SOK-222 |
Wando |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
SOK-305 |
Huksando |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
E. pungens
|
SOK-202 |
Geoje |
A |
C |
A |
C |
T |
T |
A |
G |
C |
A |
A |
A |
--- |
------ |
A |
C |
|
SOK-205 |
Geoje |
A |
C |
A |
C |
T |
T |
A |
G |
C |
A |
A |
A |
--- |
------ |
A |
C |
|
SOK-216 |
Tongyeong |
A |
C |
A |
C |
T |
T |
A |
G |
C |
A |
A |
A |
--- |
------ |
A |
C |
|
E. ×maritima
|
JJD198 |
Jeju |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
JJD243 |
Jeju |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
JJD255 |
Jeju |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
WD233 |
Wando |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
HSD252 |
Huksando |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
E. ×submacrophaylla
|
GJ376 |
Geoje |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
GJ378 |
Geoje |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
TY208 |
Tongyeong |
A |
C |
A |
C |
T |
T |
A |
G |
C |
A |
A |
A |
--- |
------ |
A |
C |
|
TY211 |
Tongyeong |
A |
C |
A |
C |
T |
T |
A |
G |
C |
A |
A |
A |
--- |
------ |
A |
C |
|
TY212 |
Tongyeong |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
|
TY213 |
Tongyeong |
C |
A |
G |
T |
A |
C |
G |
C |
C |
A |
T |
G |
AGA |
TTCTAC |
C |
G |
LITERATURE CITED
Aguilar, J. F and Feliner, G. N. 2003. Additive polymorphisms and reticulation in an ITS phylogeny of thrifts ( Armeria, Plumbaginaceae). Molecular Phylogenetics and Evolution 28: 430-447.   Aguilar, J. F. Rosselló, J. A and Feliner, G. N. 1999. Nuclear ribosomal DNA (nrDNA) concerted evolution in natural and artificial hybrids of Armeria (Plumbaginaceae). Molecular Ecology 8: 1341-1346.   Campbell, C. S. Wojciechowski, M. F. Baldwin, B. G. Alice, L. A and Donoghue, M. J. 1997. Persistent nuclear ribosomal DNA sequence polymorphism in the Amelanchier agamic complex (Rosaceae). Molecular Biology and Evolution 14: 81-90.  Cho, M.-S. Kim, C.-S. Kim, S.-H. Kim, T. O. Heo, K.-I. Jun, J and Kim, S.-C. 2014. Molecular and morphological data reveal hybrid origin of wild Prunus yedoensis (Rosaceae) from Jeju Island, Korea: Implications for the origin of the flowering cherry. American Journal of Botany 101: 1976-1986.    Du, Z.-Y. Yang, C.-F. Chen, J.-M and Guo, Y.-H. 2009. Nuclear and chloroplast DNA sequences data support the origin of Potamogeton intortusifolius J.B. He in China as a hybrid between P. perfoliatus Linn. and P. wrightii Morong. Aquatatic Botany 91: 47-50.  Falistocco, E. Passeri, V and Marconi, G. 2007. Investigations of 5S rDNA of Vitis vinifera L.: Sequence analysis and physical mapping. Genome 50: 927-938.   Gil, H.-Y and Kim, S.-C. 2016. Viola woosanensis, a recurrent spontaneous hybrid between V. ulleungdoensis and V. chaerophylloides (Violaceae) endemic to Ulleung Island, Korea. Journal of Plant Research 129: 807-822.    Karvonen, P and Savolainen, O. 1993. Variation and inheritance of ribosomal DNA in Pinus sylvestris L.(Scots pine). Heredity 71: 614-622.   Ki, K. R. 2004. Contribution of trichome morphology to the taxonomy of Korean Elaeagnus species. MS dissertation,. Sungkyunkwan University; Suwon, Korea. 43 (in Korean).
Kokubugata, G. Kurihara, T. Hirayama, Y and Obata, K. 2011. Molecular evidence for a natural hybrid origin of Ajuga ×mixta (Lamiaceae) using ITS sequence. Bullein of the National Museum of Nature and Science, Series B 37: 175-179.
Ko, S. C. 2015. Elaeagnaceae Juss. In Flora of Korea, Vol. 5b. Rosidae: Elaeagnaceae to Sapindaceae. Flora of Korea Editorial Committee (ed.), National Institute of Biological Resources, Incheon. 1-5.
Koh, J. E. 2005. A taxonomy study of the Korean Elaegnus L. (Elaeagnaceae). MS dissertation. Sungkyunkwan University; Suwon, Korea. 77 (in Korean).
Lee, T. B. 2003. Coloured Flora of Korea. Hyangmunsa, Seoul. 1: 914 Vol. 2, 910 pp. (in Korean).
Lee, W. T. 1996. Coloured Standard Illustrations of Korean Plants. Academy Publishing Co, Seoul. 624 (in Korean).
Les, D. H. Murray, N. M and Tippery, N. P. 2009. Systematics of two imperiled pondweeds ( Potamogeton vaseyi, P. gemmiparus) and taxonomic ramifications for subsection Pusilli (Potamogetonaceae). Systematic Botany 34: 643-651.  Li, W.-P. 2006. Natural hybridization between Aster ageratoides var. scaberulus and Kalimeris indica (Asteraceae): Evidence form morphology, karyotype, and ITS sequences. Botanical Studies 47: 191-197.
Liu, S.-C. Lu, C.-T and Wang, J.-C. 2009. Reticulate hybridization of Alpinia (Zingiberaceae) in Taiwan. Journal of Plant Research 122: 305-316.    Nakai, T. 1918. Notulae ad plantas Japoniae et Koreae. 18. Botanical Magazine, Tokyo. 32: 215-232.
Nakai, T. 1928. Flora Sylvatica Koreana, Vol. 17. The Forestal Experiment Station Government General of Chosen, Seoul. 1-19.
Nguyen, L.-T. Schmidt, H. A. von Haeseler, A and Minh, B. Q. 2015. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Molecular Biology and Evolution 32: 268-274.   Nieto Feliner, G. Gutiérrez Larena, B and Fuertes Aguilar, J. 2004. Fine-scale geographical structure, intra-individual polymorphism and recombination in nuclear ribosomal internal transcribed spacers in Armeria (Plumbaginaceae). Annals of Botany 93: 189-200.    Ohashi, H. 2003. Notes on Elaeagnus ×submacrophylla Servett. (Elaeagnaceae). The Journal of Japanese Botany 78: 304-307 (in Japanese).
Ohashi, H and Yoshida, S. 2003. The northernmost locality of Elaeagnus ×submacrophylla Servett. (Elaeagnaceae). The Journal of Japanese Botany 78: 301-304 (in Japanese).
Ohba. 1999. Elaeagnaceae. Flora of Japan. Vol. II c. Iwatuki, K. Boufford, D. E. Ohba, H (eds.), Kodansha, Tokyo. 152-158.
Qin, H. N and Gilbert, G. M. 2007. Elaeagnus L. In Flora of China. Vol. 13. Clusiaceae through Araliaceae. Wu, Z. Y. Raven, P. H. Hong, D. Y (eds.), Science Press, Beijing and Missouri Botanical Garden Press, St. Louis, MO. 251-270.
Rieseberg, L. H. Ellstrand, N. C and Arnold, M. 1993. What can molecular and morphological markers tell us about plant hybridization? Critical Reviews in Plant Sciences 12: 213-241.  Ronquist, F. Teslenko, M. van der Mark, P. Ayres, D. L. Darling, A. Höhna, S. Larget, B. Liu, L. Suchard, M. A and Huelsenbeck, J. P. 2012. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biology 61: 539-542.    Sang, T. Crawford, D. J and Stuessy, T. F. 1995. Documentation of reticulate evolution in peonies ( Paeonia) using internal transcribed spacer sequences of nuclear ribosomal DNA: Implications for biogeography and concerted evolution. Proceedings of the National Academy of Sciences of the United States of America 92: 6813-6817.    Schwarz, G. 1978. Estimating the dimension of a model. The Annals of Statistics 6: 461-464.  Schwarzbach, A. E and Rieseberg, L. H. 2002. Likely multiple origin of a diploid hybrid sunflower species. Molecular Ecology 11: 1703-1715.  Servettaz, C. 1909. Monographie des Elaeagnacees. Beihefte zum Botanischen Centralbatt 25: 1-420.
Shin, H. Oh, S.-H. Lim, Y. Hyun, C.-W. Cho, S.-H. Kim, Y.-I and Kim, Y.-D. 2014. Molecular evidence for hybrid origin of Aster chusanensis, an endemic species of Ulleungdo, Korea. Journal of Plant Biology 57: 174-185.   Siripun, K. C and Schilling, E. E. 2005. Molecular confirmation of the hybrid origin of Eupatorium godfreyanum (Asteraceae). American Journal of Botany 93: 319-325.  Soley, N. M and Sipes, S. D. 2021. Reproductive biology and pollinators of the invasive shrub Autumn olive (Elaeagnus umbellata Thunberg). Plant Species Biology 36: 170-186.
Stephens, M and Scheet, P. 2005. Accounting for decay of linkage disequilibrium in haplotype inference and missing-data imputation. The American Journal of Human Genetics 76: 449-462.   Stephens, M. Smith, N. J and Donnelly, P. 2001. A new statistical method for haplotype reconstruction from population data. The American Journal of Human Genetics 68: 978-989.   Suh, Y. Thien, L. B. Reeve, H. E and Zimmer, E. A. 1993. Molecular evolution and phylogenetic implications of internal transcribed spacer sequences of ribosomal DNA in Winteraceae. American Journal of Botany 80: 1042-1055.   Wendel, J.F. Schnabel, A and Seelanan, T. 1995. Bidirectional interlocus concerted evolution following allopolyploid speciation in cotton ( Gossypium). Proceedings of the National Academy of Sciences of the United States of America 92: 280-284.    White, T. J. Bruns, T. Lee, S and Taylor, J. 1990. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols: A Guide to Methods and Applications. Innis, M. A. Gelfand, D. H. Sninsky, J. J. White, T. J (eds.), Academic Press, New York. 315-322.  Wojciechowski, M. F. Lavin, M and Sanderson, M. J. 2004. A phylogeny of legumes (Leguminosae) based on analysis of the plastid mat K gene resolves many well-supported subclades within the family. American Journal of Botany 91: 1846-1862.   
|
|




















